---
title: "Release CalCOFI Database"
calcofi:
target_name: release_database
workflow_type: release
dependency:
- auto
output: data/releases
execute:
echo: true
message: true
warning: true
editor_options:
chunk_output_type: console
format:
html:
code-fold: true
editor:
markdown:
wrap: 72
---
## Overview {.unnumbered}
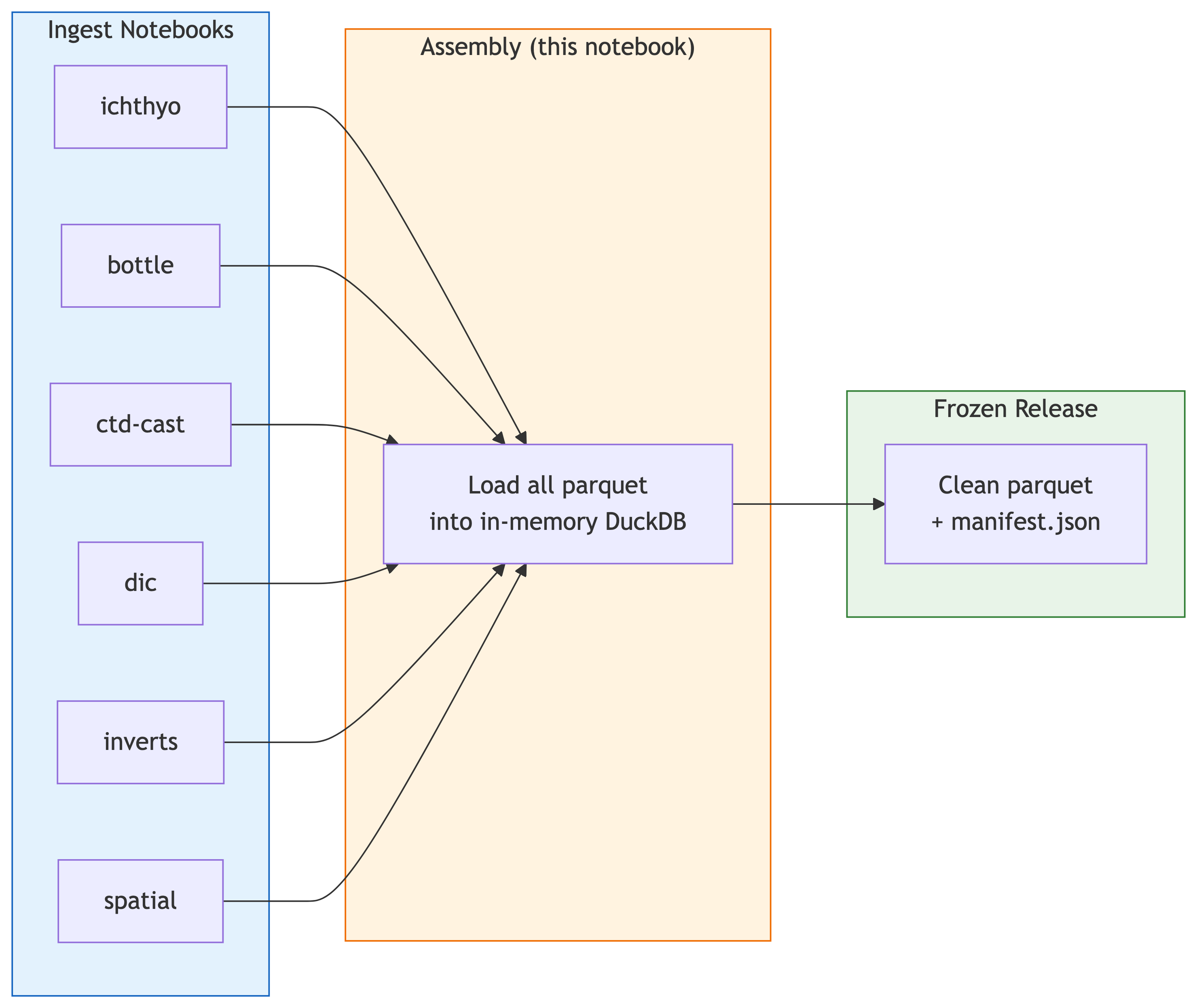
**Goal**: Create a frozen (immutable) release of the CalCOFI integrated
database by assembling all ingest parquet outputs. This is the "caboose"
notebook that always runs last, after all ingest notebooks complete.
**Upstream notebooks** are auto-discovered from `calcofi:` YAML
frontmatter in each `.qmd`. All workflows with `workflow_type: ingest`
or `spatial` feed into this release notebook via `dependency: [auto]`
in `_targets.R`.
```{mermaid}
%%| label: fig-workflow
%%| fig-cap: "Pipeline: ingest notebooks produce parquet → release caboose assembles frozen release"
flowchart LR
subgraph ingest["Ingest Notebooks"]
i1["ichthyo"]
i2["bottle"]
i3["ctd-cast"]
i4["dic"]
i5["inverts"]
i6["spatial"]
end
subgraph assembly["Assembly (this notebook)"]
a["Load all parquet<br/>into in-memory DuckDB"]
end
subgraph release["Frozen Release"]
f["Clean parquet<br/>+ manifest.json"]
end
i1 --> a
i2 --> a
i3 --> a
i4 --> a
i5 --> a
i6 --> a
a --> f
style ingest fill:#e3f2fd,stroke:#1565c0
style assembly fill:#fff3e0,stroke:#ef6c00
style release fill:#e8f4e8,stroke:#2e7d32
```
## Setup
```{r}
#| label: setup
devtools::load_all(here::here("../calcofi4db"))
devtools::load_all(here::here("../calcofi4r"))
# cleanup_gcs_obsolete(dry_run = F)
librarian::shelf(
CalCOFI / calcofi4db,
CalCOFI / calcofi4r,
DBI,
dplyr,
DT,
fs,
glue,
here,
jsonlite,
purrr,
tibble,
quiet = T
)
options(DT.options = list(scrollX = TRUE))
# release version
release_version <- format(Sys.Date(), "v%Y.%m.%d")
message(glue("Release version: {release_version}"))
```
## Assemble from Ingest Outputs
Create VIEWs on local parquet files from each ingest (zero-copy).
For tables appearing in multiple ingests, use the canonical (first) source.
```{r}
#| label: assemble_working
con_wdl <- get_duckdb_con(":memory:")
load_duckdb_extension(con_wdl, "spatial")
# auto-discover table registry from all ingest manifests
registry <- build_release_table_registry(here())
# use only canonical, non-supplemental tables
reg_canon <- registry |>
filter(canonical, !supplemental)
message(glue(
"{nrow(reg_canon)} canonical tables from ",
"{length(unique(reg_canon$ingest))} ingests"))
# create VIEWs on local parquet for each canonical table
# _new delta tables handled separately for merging
all_geom_tables <- c("grid", "site", "segment", "casts", "ctd_cast", "_spatial")
main_tables <- reg_canon |> filter(!grepl("_new$", table))
new_tables <- registry |> filter(grepl("_new$", table))
load_stats <- purrr::map_dfr(
split(main_tables, seq_len(nrow(main_tables))),
function(row) {
load_prior_tables(
con = con_wdl,
parquet_dir = row$parquet_dir,
tables = row$table,
geom_tables = all_geom_tables,
as_view = TRUE
)
})
# merge {table}_new additions into their base tables
# driven by calcofi.modifies in YAML frontmatter
if (nrow(new_tables) > 0) {
# group _new tables by their base table
base_names <- unique(sub("_new$", "", new_tables$table))
for (base_tbl in base_names) {
delta_rows <- new_tables |> filter(table == paste0(base_tbl, "_new"))
# replace VIEW with TABLE for this base table (so we can INSERT)
base_src <- main_tables |> filter(table == base_tbl)
if (nrow(base_src) > 0) {
dbExecute(con_wdl, glue("DROP VIEW IF EXISTS {base_tbl}"))
load_prior_tables(
con = con_wdl, parquet_dir = base_src$parquet_dir[1],
tables = base_tbl, geom_tables = all_geom_tables)
# get PK column for dedup
pk_col <- dbGetQuery(con_wdl, glue(
"SELECT column_name FROM information_schema.columns
WHERE table_name = '{base_tbl}'
ORDER BY ordinal_position LIMIT 1"))$column_name
for (j in seq_len(nrow(delta_rows))) {
dr <- delta_rows[j, ]
pq_path <- file.path(dr$parquet_dir, paste0(base_tbl, "_new.parquet"))
if (file.exists(pq_path)) {
dbExecute(con_wdl, glue(
"INSERT INTO {base_tbl}
SELECT * FROM read_parquet('{pq_path}')
WHERE {pk_col} NOT IN (SELECT {pk_col} FROM {base_tbl})"))
n_new <- dbGetQuery(con_wdl, glue(
"SELECT COUNT(*) AS n FROM read_parquet('{pq_path}')"))$n
message(glue("Merged {n_new} {base_tbl} addition(s) from {dr$ingest}"))
}
}
}
}
}
load_stats |>
datatable(caption = "Assembled tables (VIEWs on local parquet)")
```
## Scan Manifests for Mismatches
```{r}
#| label: scan_mismatches
# scan all ingest manifests for unresolved mismatches
all_manifests <- list.files(
"data/parquet", "manifest.json",
recursive = TRUE, full.names = TRUE)
all_mismatches <- purrr::compact(lapply(all_manifests, function(mf) {
m <- jsonlite::read_json(mf)
if (is.null(m$mismatches)) return(NULL)
dataset <- basename(dirname(mf))
purrr::imap_dfr(m$mismatches, function(items, category) {
if (length(items) == 0) return(NULL)
purrr::map_dfr(items, function(x) {
# replace NULL values with NA so as_tibble works
x[vapply(x, is.null, logical(1))] <- NA
as_tibble(x)
}) |>
mutate(dataset = dataset, category = category, .before = 1)
})
}))
if (length(all_mismatches) > 0) {
d_mismatches <- bind_rows(all_mismatches)
message(glue("{nrow(d_mismatches)} unresolved mismatch(es) across manifests"))
d_mismatches |>
datatable(caption = "Unresolved mismatches (from manifest.json)")
} else {
message("No unresolved mismatches found across manifests")
}
```
## Validate
Cross-dataset validation to ensure data integrity before freezing.
```{r}
#| label: validate
# grid_key integrity: casts.grid_key should all be in grid.grid_key
tbls <- DBI::dbListTables(con_wdl)
if (all(c("casts", "grid") %in% tbls)) {
# use information_schema to check columns (avoids GEOMETRY type issues)
casts_cols_wdl <- dbGetQuery(
con_wdl,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'casts'"
)$column_name
grid_cols_wdl <- dbGetQuery(
con_wdl,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'grid'"
)$column_name
if ("grid_key" %in% casts_cols_wdl && "grid_key" %in% grid_cols_wdl) {
grid_orphans <- dbGetQuery(
con_wdl,
"SELECT COUNT(*) AS n FROM casts c
WHERE c.grid_key IS NOT NULL
AND c.grid_key NOT IN (SELECT grid_key FROM grid)"
)$n
message(glue("Grid key orphans in casts: {grid_orphans}"))
# Grid key orphans in casts: 0
}
}
# ship PK uniqueness
if ("ship" %in% tbls) {
ship_dups <- dbGetQuery(
con_wdl,
"SELECT ship_key, COUNT(*) AS n FROM ship
GROUP BY ship_key HAVING COUNT(*) > 1"
)
if (nrow(ship_dups) > 0) {
warning(glue("Duplicate ship_key values: {nrow(ship_dups)}"))
} else {
message("ship_key: all unique")
}
}
# ship_key: all unique
# cruise PK uniqueness
if ("cruise" %in% tbls) {
cruise_dups <- dbGetQuery(
con_wdl,
"SELECT cruise_key, COUNT(*) AS n FROM cruise
GROUP BY cruise_key HAVING COUNT(*) > 1"
)
if (nrow(cruise_dups) > 0) {
warning(glue("Duplicate cruise_key values: {nrow(cruise_dups)}"))
} else {
message("cruise_key: all unique")
}
}
# cruise_key: all unique
# cruise bridge coverage
if ("casts" %in% tbls) {
bridge_stats <- dbGetQuery(
con_wdl,
"SELECT
COUNT(*) AS total_casts,
SUM(CASE WHEN ship_key IS NOT NULL THEN 1 ELSE 0 END) AS with_ship_key,
SUM(CASE WHEN cruise_key IS NOT NULL THEN 1 ELSE 0 END) AS with_cruise_key
FROM casts"
)
bridge_stats |> datatable(caption = "Cruise bridge coverage")
}
# cruise_key format validation (YYYY-MM-NODC)
if ("cruise" %in% tbls) {
bad_ck <- dbGetQuery(
con_wdl,
"SELECT cruise_key FROM cruise
WHERE cruise_key IS NOT NULL
AND NOT regexp_matches(cruise_key, '^\\d{4}-\\d{2}-.+$')"
)
if (nrow(bad_ck) > 0) {
warning(glue("cruise_key format violations: {nrow(bad_ck)} rows"))
} else {
message("cruise_key: all match YYYY-MM-NODC format")
}
}
# Warning message: cruise_key format violations: 1 rows
# cruise_key: 2019-07-
# site_key format validation (NNN.N NNN.N)
for (tbl_name in intersect(c("site", "casts", "ctd_cast"), tbls)) {
tbl_cols <- dbGetQuery(
con_wdl,
glue(
"SELECT column_name FROM information_schema.columns
WHERE table_name = '{tbl_name}'"
)
)$column_name
if ("site_key" %in% tbl_cols) {
bad_sk <- dbGetQuery(
con_wdl,
glue(
"SELECT COUNT(*) AS n FROM {tbl_name}
WHERE site_key IS NOT NULL
AND NOT regexp_matches(site_key, '^\\d{{3}}\\.\\d \\d{{3}}\\.\\d$')"
)
)$n
if (bad_sk > 0) {
warning(glue("site_key format violations in {tbl_name}: {bad_sk} rows"))
} else {
message(glue("site_key in {tbl_name}: all match NNN.N NNN.N format"))
}
}
}
# site_key in casts: all match NNN.N NNN.N format
# Warning message: site_key format violations in site: 982 rows
# cruise_summary table for Shiny app
if (all(c("cruise", "ship") %in% tbls)) {
# detect site_key vs sta_key for backward compat with old parquets
ctd_site_col <- ifelse(
"site_key" %in%
dbGetQuery(
con_wdl,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'ctd_cast'"
)$column_name,
"site_key",
"sta_key"
)
dbExecute(
con_wdl,
glue(
"
CREATE OR REPLACE TABLE cruise_summary AS
SELECT
cr.cruise_key,
EXTRACT(YEAR FROM cr.date_ym)::INTEGER AS year,
EXTRACT(MONTH FROM cr.date_ym)::INTEGER AS month,
s.ship_name,
s.ship_nodc,
COALESCE((SELECT COUNT(DISTINCT site_key) FROM site
WHERE cruise_key = cr.cruise_key), 0) AS ichthyo,
COALESCE((SELECT COUNT(DISTINCT site_key) FROM casts
WHERE cruise_key = cr.cruise_key), 0) AS bottle,
COALESCE((SELECT COUNT(DISTINCT {ctd_site_col}) FROM ctd_cast
WHERE cruise_key = cr.cruise_key), 0) AS ctd_cast,
COALESCE((SELECT COUNT(DISTINCT ds.site_key)
FROM dic_sample ds
JOIN casts c ON ds.cast_id = c.cast_id
WHERE c.cruise_key = cr.cruise_key), 0) AS dic
FROM cruise cr
JOIN ship s ON cr.ship_key = s.ship_key
ORDER BY year DESC, month DESC"
)
)
n_cs <- dbGetQuery(con_wdl, "SELECT COUNT(*) AS n FROM cruise_summary")$n
message(glue("Created cruise_summary table: {n_cs} rows"))
}
# Created cruise_summary table: 691 rows
tbl(con_wdl, "cruise_summary") |>
collect() |>
datatable(caption = "cruise_summary preview")
# run standard release validation (wrapped in tryCatch for GEOMETRY compat)
tryCatch(
{
validation <- validate_for_release(con_wdl)
if (validation$passed) {
message("Release validation passed!")
} else {
cat("Validation FAILED:\n")
cat(paste("-", validation$errors, collapse = "\n"))
}
},
error = function(e) {
message(glue("validate_for_release skipped: {e$message}"))
}
)
```
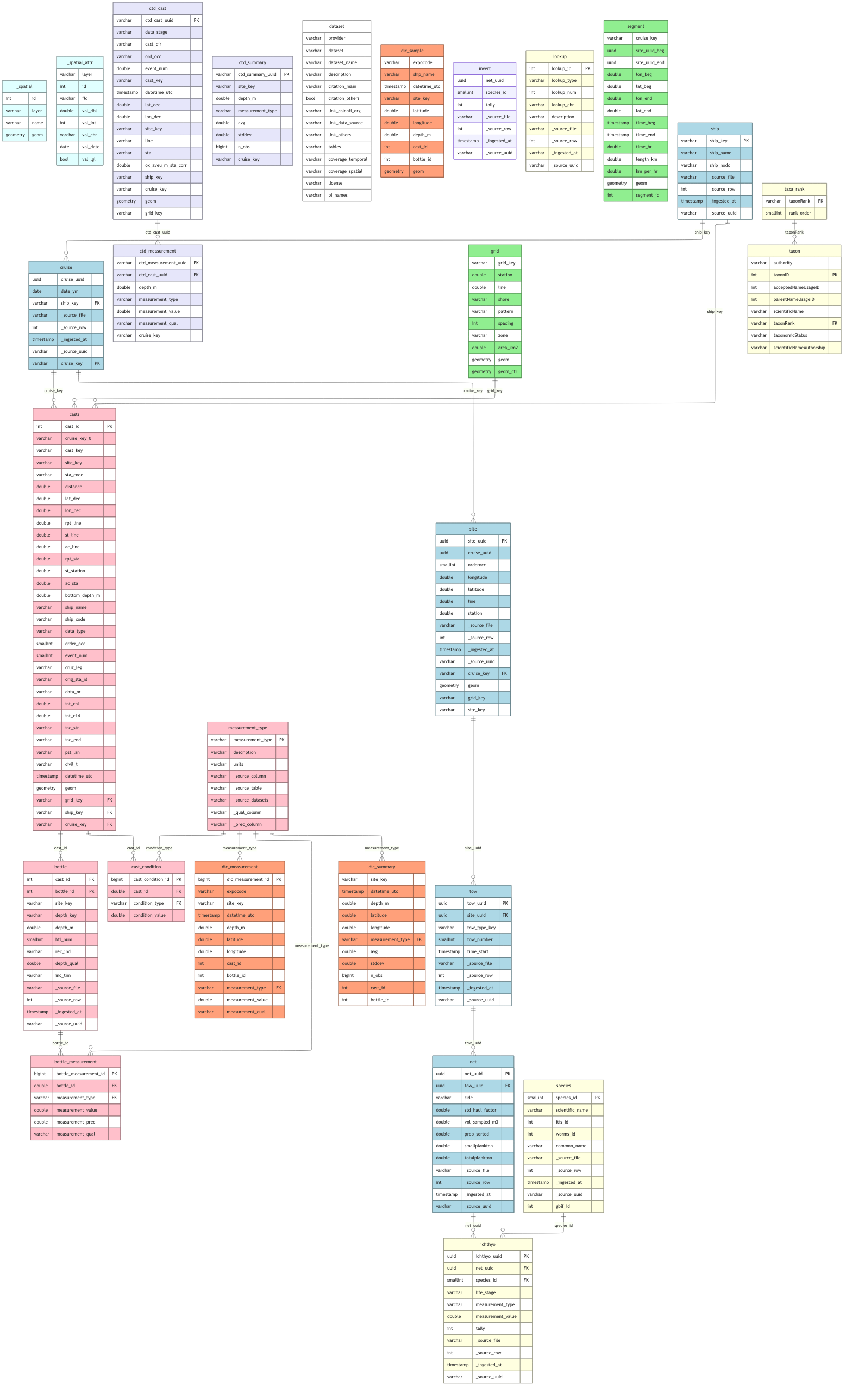
## Show Combined Schema
```{r}
# dir_frozen used later; define early so ERD can reference it
dir_frozen <- here(glue("data/releases/{release_version}"))
dir.create(dir_frozen, recursive = TRUE, showWarnings = FALSE)
erd <- cc_erd(con_wdl, layout = "elk")
plot(erd)
erd <- cc_erd(
con_wdl,
colors = list(
lightblue = c("cruise", "ship", "site", "tow", "net"),
lightyellow = c("ichthyo", "species", "lookup", "taxon", "taxa_rank"),
lightgreen = c("grid", "segment"),
pink = c(
"casts",
"bottle",
"bottle_measurement",
"cast_condition",
"measurement_type"
),
lavender = c("ctd_cast", "ctd_measurement", "ctd_summary"),
lightsalmon = c("dic_sample", "dic_measurement", "dic_summary"),
white = c("dataset")
)
)
plot(erd)
```
```{r}
#| label: combined_schema
#| fig-width: 12
#| fig-height: 10
# exclude internal tables and views
schema_tbls <- setdiff(
tbls,
c("_meta", "_sp_update", "casts_derived", "ctd_cast_derived"))
# merge per-dataset relationships.json files
rels_paths <- c(
here("data/parquet/swfsc_ichthyo/relationships.json"),
here("data/parquet/calcofi_bottle/relationships.json"),
here("data/parquet/calcofi_ctd-cast/relationships.json"),
here("data/parquet/calcofi_dic/relationships.json")
)
rels_paths <- rels_paths[file.exists(rels_paths)]
dir_frozen <- here(glue("data/releases/{release_version}"))
dir.create(dir_frozen, recursive = TRUE, showWarnings = FALSE)
rels_merged_path <- file.path(dir_frozen, "relationships.json")
if (length(rels_paths) > 0) {
merge_relationships_json(rels_paths, rels_merged_path)
# add cross-dataset FKs (bottle casts → ichthyo cruise/ship/grid)
rels_merged <- jsonlite::fromJSON(
rels_merged_path, simplifyVector = FALSE)
cross_fks <- list(
list(table = "casts", column = "cruise_key",
ref_table = "cruise", ref_column = "cruise_key"),
list(table = "casts", column = "ship_key",
ref_table = "ship", ref_column = "ship_key"),
list(table = "casts", column = "grid_key",
ref_table = "grid", ref_column = "grid_key"))
rels_merged$foreign_keys <- c(
rels_merged$foreign_keys, cross_fks)
jsonlite::write_json(
rels_merged, rels_merged_path,
auto_unbox = TRUE, pretty = TRUE, null = "null")
}
# render color-coded ERD (cc_erd handles GEOMETRY columns natively)
cc_erd(
con_wdl,
tables = schema_tbls,
rels_path = rels_merged_path,
colors = list(
lightblue = c("cruise", "ship", "site", "tow", "net"),
lightyellow = c("ichthyo", "species", "lookup",
"taxon", "taxa_rank"),
lightgreen = c("grid", "segment"),
pink = c("casts", "bottle", "bottle_measurement",
"cast_condition", "measurement_type"),
lavender = c("ctd_cast", "ctd_measurement", "ctd_summary"),
lightsalmon = c("dic_sample", "dic_measurement",
"dic_summary"),
lightcyan = c("_spatial", "_spatial_attr"),
white = "dataset"))
```
## Create Frozen Release
Strip provenance columns and export clean parquet files for public
access. See
[Frozen DuckLake](https://ducklake.select/2025/10/24/frozen-ducklake/)
pattern.
```{r}
#| label: freeze_release
dir_frozen <- here(glue("data/releases/{release_version}"))
dir_frozen_pq <- file.path(dir_frozen, "parquet")
dir.create(dir_frozen_pq, recursive = TRUE, showWarnings = FALSE)
message(glue("Creating frozen release: {release_version}"))
# only cruise_summary is derived in this notebook — export locally
# all other tables are GCS-copied from ingest/ (including provenance columns)
derived_tables <- "cruise_summary"
if (nrow(new_tables) > 0) {
# tables with _new additions need local merge + export
merged_base <- unique(sub("_new$", "", new_tables$table))
derived_tables <- c(derived_tables, merged_base)
}
# export only derived/merged tables to local parquet
export_parquet(con_wdl, "cruise_summary",
file.path(dir_frozen_pq, "cruise_summary.parquet"), compression = "zstd")
message("Exported cruise_summary.parquet")
# export merged tables (e.g., ship with _new additions)
for (tbl in setdiff(derived_tables, "cruise_summary")) {
export_parquet(con_wdl, tbl,
file.path(dir_frozen_pq, paste0(tbl, ".parquet")), compression = "zstd")
message(glue("Exported {tbl}.parquet (merged)"))
}
# build freeze stats from registry (auto-discovered)
# exclude _new delta tables (intermediate) and supplemental
freeze_stats <- reg_canon |>
filter(!supplemental, !grepl("_new$", table)) |>
select(table, rows, partitioned, gcs_prefix)
# merged tables (from _new additions) → mark as derived (gcs_prefix = NA → upload from local)
if (nrow(new_tables) > 0) {
merged_base <- unique(sub("_new$", "", new_tables$table))
freeze_stats <- freeze_stats |>
mutate(gcs_prefix = if_else(table %in% merged_base, NA_character_, gcs_prefix))
}
# add derived tables (cruise_summary, etc.)
for (dt in derived_tables) {
if (!dt %in% freeze_stats$table) {
n <- dbGetQuery(con_wdl, glue("SELECT COUNT(*) AS n FROM {dt}"))$n
freeze_stats <- freeze_stats |>
bind_rows(tibble(
table = dt, rows = n, partitioned = FALSE, gcs_prefix = NA_character_))
}
}
freeze_stats |>
datatable(caption = glue("Frozen release {release_version} — {nrow(freeze_stats)} tables"))
```
## Release Notes
```{r}
#| label: release_notes
#| results: asis
# build release notes
tables_list <- paste0(
"- ",
freeze_stats$table,
" (",
format(freeze_stats$rows, big.mark = ","),
" rows)"
)
release_notes <- paste0(
"# CalCOFI Database Release ",
release_version,
"\n\n",
"**Release Date**: ",
Sys.Date(),
"\n\n",
"## Tables Included\n\n",
paste(tables_list, collapse = "\n"),
"\n\n",
"## Total\n\n",
"- **Tables**: ",
nrow(freeze_stats),
"\n",
"- **Total Rows**: ",
format(sum(freeze_stats$rows, na.rm = TRUE), big.mark = ","),
"\n\n",
"## Data Sources\n\n",
"- `ingest_swfsc_ichthyo.qmd` - Ichthyo tables (cruise, ship, site, tow, net, species, ichthyo, grid, segment, lookup, taxon, taxa_rank)\n",
"- `ingest_calcofi_bottle.qmd` - Bottle/cast tables (casts, bottle, bottle_measurement, cast_condition, measurement_type)\n",
"- `ingest_calcofi_ctd-cast.qmd` - CTD tables (ctd_cast, ctd_measurement, ctd_summary, measurement_type)\n",
"- `ingest_calcofi_dic.qmd` - DIC/alkalinity tables (dic_sample, dic_measurement, dic_summary, dataset)\n\n",
"## Cross-Dataset Integration\n\n",
"- **Ship matching**: Reconciled ship codes between bottle casts and swfsc ship reference\n",
"- **Cruise bridge**: Derived cruise_key (YYYY-MM-NODC) for bottle casts via ship matching + datetime\n",
"- **Taxonomy**: Standardized species with WoRMS AphiaID, ITIS TSN, GBIF backbone key\n",
"- **Taxon hierarchy**: Built taxon + taxa_rank tables from WoRMS/ITIS classification\n\n",
"## Access\n\n",
"Parquet files can be queried directly from GCS:\n\n",
"```r\n",
"library(duckdb)\n",
"con <- dbConnect(duckdb())\n",
"dbExecute(con, 'INSTALL httpfs; LOAD httpfs;')\n",
"dbGetQuery(con, \"\n",
" SELECT * FROM read_parquet(\n",
" 'https://storage.googleapis.com/calcofi-db/ducklake/releases/",
release_version,
"/parquet/ichthyo.parquet')\n",
" LIMIT 10\")\n",
"```\n\n",
"Or use calcofi4r:\n\n",
"```r\n",
"library(calcofi4r)\n",
"con <- cc_get_db(version = '",
release_version,
"')\n",
"```\n"
)
writeLines(release_notes, file.path(dir_frozen, "RELEASE_NOTES.md"))
message(glue(
"Release notes written to {file.path(dir_frozen, 'RELEASE_NOTES.md')}"
))
cat(release_notes)
```
## Upload Frozen Release to GCS
```{r}
#| label: upload_frozen
gcs_bucket <- "calcofi-db"
gcs_release <- glue("ducklake/releases/{release_version}")
gcloud <- find_gcloud()
# 1. GCS server-side copy for ingest tables (auto-discovered from registry)
copy_rows <- freeze_stats |> filter(!is.na(gcs_prefix))
message(glue("Copying {nrow(copy_rows)} tables from ingest/ to releases/ on GCS..."))
for (i in seq_len(nrow(copy_rows))) {
tbl <- copy_rows$table[i]
pfx <- copy_rows$gcs_prefix[i]
part <- copy_rows$partitioned[i]
if (part) {
# partitioned: copy directory
src <- glue("gs://{gcs_bucket}/{pfx}/{tbl}")
dst <- glue("gs://{gcs_bucket}/{gcs_release}/parquet/{tbl}")
res <- system2(gcloud, c("storage", "cp", "-r",
paste0(src, "/*"), dst), stdout = TRUE, stderr = TRUE)
} else {
src <- glue("gs://{gcs_bucket}/{pfx}/{tbl}.parquet")
dst <- glue("gs://{gcs_bucket}/{gcs_release}/parquet/{tbl}.parquet")
res <- system2(gcloud, c("storage", "cp",
src, dst), stdout = TRUE, stderr = TRUE)
}
rc <- attr(res, "status") %||% 0L
if (rc != 0) {
stop(glue("GCS copy failed for {tbl}: {src} -> {dst}\n",
" exit code {rc}: {paste(res, collapse = '\n')}"))
}
message(glue(" {tbl}: copied from {pfx}"))
}
# 2. upload derived tables from local parquet (cruise_summary)
derived_local <- list.files(dir_frozen_pq, pattern = "[.]parquet$",
full.names = TRUE)
for (pq in derived_local) {
tbl <- tools::file_path_sans_ext(basename(pq))
gcs_path <- glue("gs://{gcs_bucket}/{gcs_release}/parquet/{tbl}.parquet")
put_gcs_file(pq, gcs_path)
message(glue(" {tbl}: uploaded (derived)"))
}
# 3. build and upload catalog.json (needed by cc_get_db())
tables_df <- freeze_stats |>
select(name = table, rows, partitioned)
catalog <- list(
version = release_version,
release_date = as.character(Sys.Date()),
total_rows = sum(tables_df$rows, na.rm = TRUE),
total_size = 0,
tables = tables_df)
catalog_path <- file.path(dir_frozen, "catalog.json")
jsonlite::write_json(catalog, catalog_path, auto_unbox = TRUE, pretty = TRUE)
put_gcs_file(catalog_path,
glue("gs://{gcs_bucket}/{gcs_release}/catalog.json"))
# upload RELEASE_NOTES.md
notes_path <- file.path(dir_frozen, "RELEASE_NOTES.md")
if (file.exists(notes_path))
put_gcs_file(notes_path,
glue("gs://{gcs_bucket}/{gcs_release}/RELEASE_NOTES.md"))
# upload relationships.json
rels_json <- file.path(dir_frozen, "relationships.json")
if (file.exists(rels_json))
put_gcs_file(rels_json,
glue("gs://{gcs_bucket}/{gcs_release}/relationships.json"))
# 4. update versions.json and latest.txt
# discover all releases from GCS and rebuild versions.json
gcs_ls <- system2(gcloud, c("storage", "ls",
glue("gs://{gcs_bucket}/ducklake/releases/")),
stdout = TRUE, stderr = TRUE)
release_vers <- regmatches(gcs_ls,
regexpr("v[0-9]{4}[.][0-9]{2}[.]*[0-9]*", gcs_ls))
https_base <- glue("https://storage.googleapis.com/{gcs_bucket}/ducklake/releases")
all_versions <- purrr::compact(lapply(release_vers, function(v) {
tryCatch({
cat_data <- jsonlite::fromJSON(glue("{https_base}/{v}/catalog.json"))
list(
version = cat_data$version,
release_date = cat_data$release_date %||% NA_character_,
tables = if (is.data.frame(cat_data$tables)) nrow(cat_data$tables)
else length(cat_data$tables),
total_rows = as.integer(cat_data$total_rows %||% 0),
size_mb = round((cat_data$total_size %||% 0) / 1024 / 1024, 1))
}, error = function(e) NULL)
}))
all_versions <- all_versions[order(
sapply(all_versions, `[[`, "version"), decreasing = TRUE)]
versions_local <- tempfile(fileext = ".json")
jsonlite::write_json(list(versions = all_versions), versions_local,
auto_unbox = TRUE, pretty = TRUE)
put_gcs_file(versions_local,
glue("gs://{gcs_bucket}/ducklake/releases/versions.json"))
# update latest.txt
latest_local <- tempfile()
writeLines(release_version, latest_local)
put_gcs_file(latest_local,
glue("gs://{gcs_bucket}/ducklake/releases/latest.txt"))
message(glue(
"Release {release_version} published ({length(all_versions)} versions tracked)"))
```
## Cleanup
```{r}
#| label: cleanup
# close in-memory DuckDB connection
close_duckdb(con_wdl)
message("Assembly DuckDB connection closed")
# summary
message(glue("\n=== Summary ==="))
message(glue("Frozen release: {release_version} created at {dir_frozen}"))
message(glue("Tables: {nrow(freeze_stats)}"))
message(glue("Total rows: {format(sum(freeze_stats$rows, na.rm = TRUE), big.mark = ',')}"))
```
::: {.callout-caution collapse="true"}
## Session Info
```{r session_info}
devtools::session_info()
```
:::