---
title: "Ingest Spatial Boundaries"
calcofi:
target_name: ingest_spatial
workflow_type: spatial
dependency: []
output: data/parquet/spatial/manifest.json
editor_options:
chunk_output_type: console
---
## Overview
Process spatial boundary datasets for:
1. **Map overlay** — PMTiles served via `file.calcofi.io/_spatial/`
2. **Spatial queries** — `_spatial` + `_spatial_attr` tables in CalCOFI DuckDB
3. **Future summarization** — aggregate CalCOFI data into boundary features
**Source**: `~/My Drive/projects/calcofi/data-public/_spatial/`
([US Maritime Limits](https://nauticalcharts.noaa.gov/data/us-maritime-limits-and-boundaries.html),
CA CDFW, BOEM, NOAA ONMS, MEOW, and others)
**Registry**: `metadata/spatial_layers.csv` — one row per visual layer, with
dataset grouping, styling, filter expressions, and source file paths.
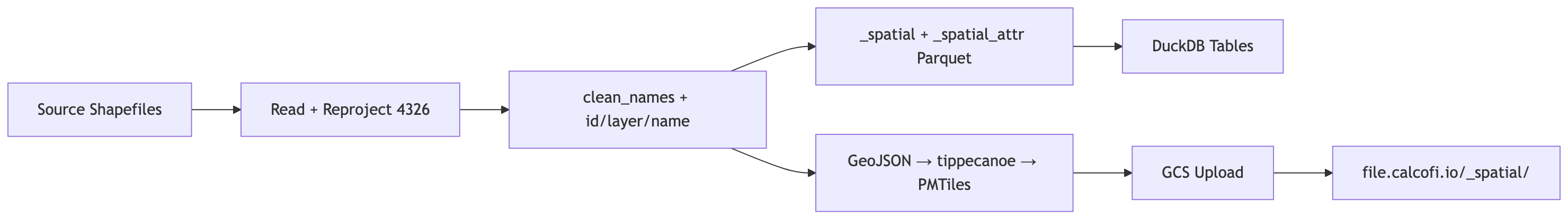
### Data Flow
```{mermaid}
graph LR
A[Source Shapefiles] --> B[Read + Reproject 4326]
B --> C["clean_names + id/layer/name"]
C --> D["_spatial + _spatial_attr Parquet"]
C --> E["GeoJSON → tippecanoe → PMTiles"]
D --> F[DuckDB Tables]
E --> G[GCS Upload]
G --> H["file.calcofi.io/_spatial/"]
```
### Pre-processing
Each dataset is standardized:
- **CRS**: reprojected to `EPSG:4326` (WGS 84 unprojected)
- **Columns**: `janitor::clean_names()` for snake_case
- **ID**: integer `id` column (per-layer sequential)
- **layer**: readable layer name from `metadata/spatial_layers.csv`
- **name**: per-feature label from `name_field` (or `layer` for dissolved layers)
## Setup
```{r}
#| label: setup
librarian::shelf(
arcgislayers,
calcofi / calcofi4db,
DBI,
dplyr,
DT,
duckdb,
fs,
glue,
here,
janitor,
jsonlite,
purrr,
readr,
sf,
stringr,
tibble,
tidyr,
quiet = T
)
# common ingest settings (overwrite, dir_data)
source(here("libs/ingest.R"))
dir_spatial <- path_expand(glue("{dir_data}/_spatial"))
dir_pmtiles <- here("data/pmtiles")
dir_parquet <- here("data/parquet/spatial")
gcs_bucket <- "calcofi-files-public"
gcs_prefix <- "_spatial"
dir_create(dir_pmtiles)
dir_create(dir_parquet)
# archive source spatial files to GCS
# gcloud config set account bebest@ucsd.edu
# TODO: fix this since it seems to put into /archive/_spatial without a timestamp
# sync_to_gcs(
# local_dir = dir_spatial,
# gcs_prefix = "archive/_spatial",
# bucket = gcs_bucket,
# exclude = c(".DS_Store", "*.tmp")
# )
# load registry
d_layers <- read_csv(
here("metadata/spatial_layers.csv"),
show_col_types = FALSE
)
datatable(
d_layers,
caption = "Spatial layers registry from `metadata/spatial_layers.csv`"
)
# compare with folders/files in dir_spatial
spatial_dirs <- dir_ls(dir_spatial, type = "directory")
spatial_dirs_not_layers_csv <- setdiff(
basename(spatial_dirs),
unique(d_layers$source_folder)
)
if (length(spatial_dirs_not_layers_csv) > 0) {
cat(
glue(
"WARNING: The following folders exist in `{dir_spatial}` but are not ",
"referenced in `metadata/spatial_layers.csv`:\n",
paste0("- ", spatial_dirs_not_layers_csv, collapse = "\n"),
"\n\n"
)
)
}
# unique dataset groups (one pmtiles per group)
d_groups <- d_layers |>
distinct(dataset_group, source_folder, source_file, source_format) |>
mutate(
source_path = path(dir_spatial, source_folder, source_file),
file_exists = file_exists(source_path)
)
cat(
glue("{nrow(d_layers)} layers across {nrow(d_groups)} dataset groups"),
"\n"
)
```
## ArcGIS Rest layer(s) to gpkg: `CA_watershed-boundaries`
```{r}
#| label: arcgis-rest
fs_to_sf <- function(url, where = NULL) {
fs <- arcgislayers::arc_open(url)
lyr <- arcgislayers::get_layer(fs, 0)
sf <- arcgislayers::arc_select(
lyr,
where = where
) |>
tibble::tibble() |>
sf::st_as_sf() |>
janitor::clean_names()
return(sf)
}
# [USGS Watershed Boundary Dataset (WBD) HUC8, California Vicinity - Overview](https://www.arcgis.com/home/item.html?id=a98ef9ead82a4bd6aa09d46869f75a49&sublayer=0)
url_rest <- "https://services2.arcgis.com/Uq9r85Potqm3MfRV/arcgis/rest/services/WBD_HU8_wm/FeatureServer"
out_gpkg <- glue("{dir_spatial}/CA_watershed-boundaries/wbd-hu8_ca.gpkg")
if (!file_exists(out_gpkg) || overwrite) {
fs_to_sf(url_rest) |>
write_sf(out_gpkg, delete_dsn = TRUE)
}
```
## Clip CA Current LME to US EEZ
```{r}
#| label: clip-layers
cacurrent_geo <- glue(
"{dir_spatial}/NOAA_IEA-CA-Current/lme-ca-current.geojson"
)
usaeez_geo <- glue(
"{dir_spatial}/NOAA_IEA-CA-Current/usa-eez.geojson"
)
cacurrent_usaeez_geo <- glue(
"{dir_spatial}/NOAA_IEA-CA-Current/lme-ca-current_usa-eez.geojson"
)
if (!file_exists(cacurrent_usaeez_geo) || overwrite) {
cacurrent <- st_read(cacurrent_geo, quiet = TRUE)
usaeez <- st_read(usaeez_geo, quiet = TRUE)
cacurrent_usaeez <- st_intersection(cacurrent, usaeez)
st_write(
cacurrent_usaeez,
cacurrent_usaeez_geo,
delete_dsn = TRUE,
quiet = TRUE
)
}
```
## Helper functions
```{r}
#| label: helpers
# read source spatial data from various formats
read_spatial_source <- function(
source_folder,
source_file,
source_format,
dir_spatial
) {
if (source_format == "rda") {
e <- new.env()
load(path(dir_spatial, source_folder, source_file), envir = e)
for (nm in ls(e)) {
if (inherits(get(nm, envir = e), "sf")) {
return(get(nm, envir = e) |> select(where(~ !is.list(.))))
}
}
return(NULL)
} else if (source_format == "gdb") {
parts <- str_split_1(source_file, "\\|")
st_read(
path(dir_spatial, source_folder, parts[1]),
layer = parts[2],
quiet = TRUE
)
} else {
st_read(
path(dir_spatial, source_folder, source_file),
quiet = TRUE
)
}
}
# apply a MapLibre filter expression as R filter on sf
# supports: ["==", ["get", "col"], val]
apply_filter_expr <- function(sf_data, filter_expr_json) {
if (is.na(filter_expr_json) || filter_expr_json == "") {
return(sf_data)
}
expr <- jsonlite::fromJSON(filter_expr_json, simplifyVector = FALSE)
if (
length(expr) == 3 &&
expr[[1]] == "==" &&
is.list(expr[[2]]) &&
expr[[2]][[1]] == "get"
) {
col_name <- expr[[2]][[2]]
val <- expr[[3]]
sf_data |> filter(.data[[col_name]] == val)
} else {
warning(glue("unsupported filter_expr: {filter_expr_json}"))
sf_data
}
}
# build _spatial_attr rows from an sf object
build_spatial_attr <- function(sf_data, layer_name) {
# exclude core columns and geometry
geom_col <- attr(sf_data, "sf_column")
core_cols <- c("id", "layer", "name", geom_col)
attr_cols <- setdiff(names(sf_data), core_cols)
if (length(attr_cols) == 0) {
return(tibble())
}
# pivot each attribute column into EAV rows
map_dfr(attr_cols, function(col) {
vals <- sf_data[[col]]
cls <- class(vals)[1]
base <- tibble(
layer = layer_name,
id = sf_data$id,
fld = col,
val_dbl = NA_real_,
val_int = NA_integer_,
val_chr = NA_character_,
val_date = as.Date(NA),
val_lgl = NA
)
if (cls %in% c("numeric", "double")) {
base$val_dbl <- as.double(vals)
} else if (cls == "integer") {
base$val_int <- as.integer(vals)
} else if (cls %in% c("character", "factor")) {
base$val_chr <- as.character(vals)
} else if (cls == "Date") {
base$val_date <- as.Date(vals)
} else if (cls == "logical") {
base$val_lgl <- as.logical(vals)
} else {
# fallback: coerce to character
base$val_chr <- as.character(vals)
}
base
})
}
```
## Process datasets
For each dataset group: read source, add `id`/`layer`/`name` columns,
generate PMTiles, and accumulate data for consolidated `_spatial` tables.
```{r}
#| label: process-datasets
#| output: asis
# accumulators for consolidated tables
all_spatial <- list()
all_spatial_attr <- list()
for (i in seq_len(nrow(d_groups))) {
grp <- d_groups[i, ]
dataset_group <- grp$dataset_group
pmtiles_path <- path(dir_pmtiles, glue("{dataset_group}.pmtiles"))
# skip if already processed and not redoing
if (!overwrite && file_exists(pmtiles_path)) {
cat(glue("- **{dataset_group}**: skipped (already exists)\n\n"))
next
}
# read source data
sf_data <- tryCatch(
read_spatial_source(
grp$source_folder,
grp$source_file,
grp$source_format,
dir_spatial
),
error = function(e) {
cat(glue(
"- **{dataset_group}**: SKIPPED — failed to read ",
"`{grp$source_folder}/{grp$source_file}`: {e$message}\n\n"
))
NULL
}
)
if (is.null(sf_data) || nrow(sf_data) == 0) {
cat(glue("- **{dataset_group}**: SKIPPED — empty dataset\n\n"))
next
}
# reproject to EPSG:4326
src_crs <- st_crs(sf_data)$epsg
if (is.na(src_crs) || src_crs != 4326) {
sf_data <- st_transform(sf_data, 4326)
}
# clean column names
sf_data <- clean_names(sf_data)
# get layer rows for this group
grp_layers <- d_layers |> filter(dataset_group == !!dataset_group)
if (nrow(grp_layers) == 1 && is.na(grp_layers$filter_expr[1])) {
# single layer, no filter
layer_name <- grp_layers$layer[1]
name_field <- grp_layers$name_field[1]
if (is.na(name_field) || name_field == "") {
# dissolve all features into one (st_make_valid for bad geometries)
sf_data <- sf_data |>
st_make_valid() |>
summarise(geometry = st_union(geometry)) |>
mutate(
id = 1L,
layer = layer_name,
name = layer_name
) |>
relocate(id, layer, name)
} else {
sf_data <- sf_data |>
mutate(
id = row_number() |> as.integer(),
layer = layer_name,
name = as.character(.data[[name_field]])
) |>
relocate(id, layer, name)
}
# accumulate for _spatial tables
geom_col_name <- attr(sf_data, "sf_column")
all_spatial[[layer_name]] <- sf_data |>
select(id, layer, name, all_of(geom_col_name))
all_spatial_attr[[layer_name]] <- build_spatial_attr(sf_data, layer_name)
} else {
# multi-layer group (e.g., noaa_maritime_boundaries with filter_exprs)
parts_list <- list()
for (j in seq_len(nrow(grp_layers))) {
lr <- grp_layers[j, ]
layer_name <- lr$layer
name_field <- lr$name_field
sf_filtered <- apply_filter_expr(sf_data, lr$filter_expr)
if (nrow(sf_filtered) == 0) {
next
}
if (is.na(name_field) || name_field == "") {
sf_filtered <- sf_filtered |>
st_make_valid() |>
summarise(geometry = st_union(geometry)) |>
mutate(id = 1L, layer = layer_name, name = layer_name) |>
relocate(id, layer, name)
} else {
sf_filtered <- sf_filtered |>
mutate(
id = row_number() |> as.integer(),
layer = layer_name,
name = as.character(.data[[name_field]])
) |>
relocate(id, layer, name)
}
parts_list[[layer_name]] <- sf_filtered
geom_col_name <- attr(sf_filtered, "sf_column")
all_spatial[[layer_name]] <- sf_filtered |>
select(id, layer, name, all_of(geom_col_name))
all_spatial_attr[[layer_name]] <- build_spatial_attr(
sf_filtered,
layer_name
)
}
# for PMTiles: use the full original sf_data with layer/name added per-feature
# tag each feature with its layer based on which filter it matches
sf_data <- sf_data |> mutate(layer = NA_character_, name = NA_character_)
for (j in seq_len(nrow(grp_layers))) {
lr <- grp_layers[j, ]
if (!is.na(lr$filter_expr) && lr$filter_expr != "") {
expr <- jsonlite::fromJSON(lr$filter_expr, simplifyVector = FALSE)
if (
length(expr) == 3 &&
expr[[1]] == "==" &&
is.list(expr[[2]]) &&
expr[[2]][[1]] == "get"
) {
col_nm <- expr[[2]][[2]]
val <- expr[[3]]
mask <- sf_data[[col_nm]] == val
mask[is.na(mask)] <- FALSE
sf_data$layer[mask] <- lr$layer
nf <- lr$name_field
if (!is.na(nf) && nf != "") {
sf_data$name[mask] <- as.character(sf_data[[nf]][mask])
} else {
sf_data$name[mask] <- lr$layer
}
}
}
}
# features not matching any filter: use group name
sf_data <- sf_data |>
mutate(
layer = if_else(is.na(layer), dataset_group, layer),
name = if_else(is.na(name), layer, name)
)
if (!"id" %in% names(sf_data)) {
sf_data <- sf_data |> mutate(id = row_number() |> as.integer())
} else {
sf_data <- sf_data |> mutate(id = as.integer(id))
}
sf_data <- sf_data |> relocate(id, layer, name)
}
# write PMTiles via tippecanoe
geojson_tmp <- tempfile(fileext = ".geojson")
st_write(sf_data, geojson_tmp, delete_dsn = TRUE, quiet = TRUE)
tippecanoe_args <- c(
"-o",
pmtiles_path,
"-z10",
"-Z0",
"--simplification=10",
"--no-tile-size-limit",
"-l",
dataset_group,
"--force",
geojson_tmp
)
system2("tippecanoe", args = tippecanoe_args, stdout = TRUE, stderr = TRUE)
unlink(geojson_tmp)
if (!file_exists(pmtiles_path)) {
cat(glue("- **{dataset_group}**: SKIPPED — tippecanoe failed\n\n"))
next
}
pmtiles_size <- file_size(pmtiles_path)
cat(glue(
"- **{dataset_group}**: {nrow(sf_data)} features ",
"(CRS {src_crs} -> 4326), PMTiles {pmtiles_size}\n\n"
))
}
```
## Build consolidated `_spatial` + `_spatial_attr` parquet
```{r}
#| label: build-spatial-parquet
pq_spatial <- path(dir_parquet, "_spatial.parquet")
pq_spatial_attr <- path(dir_parquet, "_spatial_attr.parquet")
# skip if parquet already exists and no new layers were processed
if (length(all_spatial) == 0 && file_exists(pq_spatial)) {
message("_spatial parquet already exists, skipping rebuild")
# read back for downstream references
con_tmp <- dbConnect(duckdb())
dbExecute(con_tmp, "INSTALL spatial; LOAD spatial;")
spatial_combined <- st_read(
con_tmp,
query = glue("SELECT * FROM read_parquet('{pq_spatial}')")
)
attr_combined <- arrow::read_parquet(pq_spatial_attr)
dbDisconnect(con_tmp, shutdown = TRUE)
} else {
# combine all layers into single sf
spatial_combined <- bind_rows(all_spatial) |>
st_as_sf()
message(glue(
"_spatial: {nrow(spatial_combined)} features across ",
"{length(unique(spatial_combined$layer))} layers"
))
# combine all attributes
attr_combined <- bind_rows(all_spatial_attr)
message(glue("_spatial_attr: {nrow(attr_combined)} rows"))
# remove old per-dataset parquet files
old_parquets <- dir_ls(dir_parquet, glob = "*.parquet")
old_parquets <- old_parquets[!grepl("^_spatial", basename(old_parquets))]
if (length(old_parquets) > 0) {
file_delete(old_parquets)
message(glue(
"Removed {length(old_parquets)} old per-dataset parquet files"
))
}
# write _spatial via DuckDB (preserves geometry as WKB in parquet)
spatial_geojson <- tempfile(fileext = ".geojson")
st_write(spatial_combined, spatial_geojson, delete_dsn = TRUE, quiet = TRUE)
con_tmp <- dbConnect(duckdb())
dbExecute(con_tmp, "INSTALL spatial; LOAD spatial;")
dbExecute(
con_tmp,
glue(
"COPY (SELECT * FROM ST_Read('{spatial_geojson}'))
TO '{pq_spatial}' (FORMAT PARQUET)"
)
)
dbDisconnect(con_tmp, shutdown = TRUE)
unlink(spatial_geojson)
# write _spatial_attr as plain parquet
arrow::write_parquet(attr_combined, pq_spatial_attr)
message(glue(
"Wrote _spatial.parquet ({file_size(pq_spatial)}) ",
"and _spatial_attr.parquet ({file_size(pq_spatial_attr)})"
))
} # end else (rebuild)
```
## Upload to GCS
```{r}
#| label: upload-gcs
# upload PMTiles to calcofi-files-public bucket
cat(glue(
"Uploading PMTiles to gs://{gcs_bucket}/{gcs_prefix}/\n\n"
))
sync_to_gcs(
local_dir = dir_pmtiles,
gcs_prefix = gcs_prefix,
bucket = gcs_bucket,
pattern = "\\.pmtiles$"
)
# upload parquets to calcofi-db ingest bucket (for release_database.qmd)
dir_label <- basename(dir_parquet) # "spatial"
gcs_ingest_prefix <- glue("ingest/{dir_label}")
gcs_ingest_bucket <- "calcofi-db"
cat(glue(
"\nUploading parquets to gs://{gcs_ingest_bucket}/{gcs_ingest_prefix}/\n\n"
))
sync_to_gcs(
local_dir = dir_parquet,
gcs_prefix = gcs_ingest_prefix,
bucket = gcs_ingest_bucket
)
```
## Load into DuckDB
```{r}
#| label: load-duckdb
db_path <- here("data/calcofi_wrangling.duckdb")
con <- dbConnect(duckdb(dbdir = db_path))
dbExecute(con, "INSTALL spatial; LOAD spatial;")
pq_spatial <- path(dir_parquet, "_spatial.parquet")
pq_spatial_attr <- path(dir_parquet, "_spatial_attr.parquet")
# load _spatial with EPSG:4326
dbExecute(
con,
glue(
"CREATE OR REPLACE TABLE _spatial AS
SELECT id, layer, name,
ST_GeomFromWKB(ST_AsWKB(geom)) AS geom
FROM read_parquet('{pq_spatial}')"
)
)
n_spatial <- dbGetQuery(con, "SELECT COUNT(*) AS n FROM _spatial")$n
message(glue("_spatial: {n_spatial} rows"))
# load _spatial_attr (no geometry)
dbExecute(
con,
glue(
"CREATE OR REPLACE TABLE _spatial_attr AS
SELECT * FROM read_parquet('{pq_spatial_attr}')"
)
)
n_attr <- dbGetQuery(con, "SELECT COUNT(*) AS n FROM _spatial_attr")$n
message(glue("_spatial_attr: {n_attr} rows"))
# drop old spatial_* tables if they exist
old_tbls <- dbListTables(con)
old_spatial <- old_tbls[grepl("^spatial_", old_tbls)]
for (tbl in old_spatial) {
dbExecute(con, glue("DROP TABLE IF EXISTS {tbl}"))
message(glue("Dropped old table: {tbl}"))
}
dbDisconnect(con, shutdown = TRUE)
```
## Write manifest
```{r}
#| label: manifest
write_spatial_manifest(dir_parquet)
```
## Output summary
```{r}
#| label: summary
#| output: asis
cat("### PMTiles\n\n")
dir_ls(dir_pmtiles, glob = "*.pmtiles") |>
tibble(path = _) |>
mutate(
file = path_file(path),
size = file_size(path)
) |>
select(file, size) |>
knitr::kable()
cat("\n### Consolidated Parquet\n\n")
dir_ls(dir_parquet, glob = "*.parquet") |>
tibble(path = _) |>
mutate(
file = path_file(path),
size = file_size(path)
) |>
select(file, size) |>
knitr::kable()
cat(glue(
"\n### Database Tables\n\n",
"- `_spatial`: {nrow(spatial_combined)} features, ",
"{length(unique(spatial_combined$layer))} layers\n",
"- `_spatial_attr`: {nrow(attr_combined)} attribute rows\n"
))
```