---
title: "Ingest Bottle Database"
calcofi:
target_name: ingest_calcofi_bottle
workflow_type: ingest
dependency:
- ingest_swfsc_ichthyo
output: data/parquet/calcofi_bottle/manifest.json
modifies:
- ship
---
## Overview
**Source**: [Bottle Database | CalCOFI.org](https://calcofi.org/data/oceanographic-data/bottle-database/)
- Download: <https://calcofi.org/downloads/database/CalCOFI_Database_194903-202105_csv_16October2023.zip>
**Goal**: Ingest bottle database from source CSV files into a local
wrangling DuckDB, then export as parquet files for downstream use.
**Two source tables**:
- `194903-202105_Cast.csv` (61 fields) — station-level: cruise,
location, meteorology, integrated measurements
- `194903-202105_Bottle.csv` (62 fields) — sample-level: depth,
temperature, salinity, O₂, nutrients, pigments, C14, pH
**Coverage**: 1949-03 to 2021-05 (72 years)
**Join key**: `cast_id` (in both tables; source column `Cst_Cnt`)
This workflow processes CalCOFI bottle database CSV files and outputs
parquet files. The workflow:
1. Reads CSV files from source directory (with GCS archive sync)
2. Loads into local wrangling DuckDB with column renames
3. Applies ship name/code corrections
4. Consolidates temporal columns into `datetime_utc`, drops derivable columns, creates `casts_derived` VIEW
5. Pivots bottle measurements into long-format `bottle_measurement` table
6. Pivots cast environmental conditions into `cast_condition` table
7. Loads `measurement_type` reference table
8. Validates primary keys and foreign key integrity
9. Adds spatial geometry and grid assignments
10. Exports to parquet files (for later integration into Working DuckLake)
```{r}
#| label: setup
# devtools::install_local(here::here("../calcofi4db"), force = T)
devtools::load_all(here::here("../calcofi4db"))
devtools::load_all(here::here("../calcofi4r"))
librarian::shelf(
CalCOFI / calcofi4db,
CalCOFI / calcofi4r,
DBI,
dplyr,
DT,
fs,
glue,
gargle,
googledrive,
here,
htmltools,
janitor,
jsonlite,
knitr,
listviewer,
litedown,
lubridate,
mapview,
purrr,
readr,
rlang,
sf,
stringr,
tibble,
tidyr,
units,
webshot2,
quiet = T
)
options(readr.show_col_types = F)
options(DT.options = list(scrollX = TRUE))
# define paths
# common ingest settings (overwrite, dir_data)
source(here("libs/ingest.R"))
dataset_name <- "CalCOFI Bottle Database"
provider <- "calcofi"
dataset <- "bottle"
dir_label <- glue("{provider}_{dataset}")
subdir <- "CalCOFI_Database_194903-202105_csv_16October2023"
dir_parquet <- here(glue("data/parquet/{dir_label}"))
db_path <- here(glue("data/wrangling/{dir_label}.duckdb"))
# load data using calcofi4db package
# - reads from local Google Drive mount
# - syncs to GCS archive if files changed (creates new timestamped archive)
# - tracks GCS archive path for provenance
dir_dataset <- glue("{dir_data}/{provider}/{dataset}")
# unified measurement_type reference (needed for pivot mappings)
meas_type_csv <- here("metadata/measurement_type.csv")
```
```{r}
#| label: load_db
if (overwrite) {
if (file_exists(db_path)) {
file_delete(db_path)
}
if (dir_exists(dir_parquet)) dir_delete(dir_parquet)
}
dir_create(dirname(db_path))
con <- get_duckdb_con(db_path)
load_duckdb_extension(con, "spatial")
```
```{r}
#| label: read_csv_files
d_meas_type <- read_csv(meas_type_csv)
d <- read_csv_files(
provider = provider,
dataset = dataset,
subdir = subdir,
dir_data = dir_data,
sync_archive = TRUE,
metadata_dir = here("metadata"),
field_descriptions = list(
"194903-202105_Cast" = glue(
"{dir_dataset}/Cast Field Descriptions.csv"
),
"194903-202105_Bottle" = glue(
"{dir_dataset}/Bottle Field Descriptions - UTF-8.csv"
)
)
)
# show source files summary
message(glue("Loaded {nrow(d$source_files)} tables from {d$paths$dir_csv}"))
message(glue("Total rows: {sum(d$source_files$nrow)}"))
```
## Check for any mismatched tables and fields
```{r}
#| label: data_integrity_check
# check data integrity
# type_exceptions = "all": type mismatches (34) are expected since readr
# infers types differently (e.g. quality codes as numeric, Time as hms);
# these are resolved during ingestion via flds_redefine.csv type_new
integrity <- check_data_integrity(
d = d,
dataset_name = dataset_name,
type_exceptions = "all"
)
```
```{r}
#| label: data_integrity_message
#| output: asis
#| echo: false
# render the pass/fail message as markdown
render_integrity_message(integrity)
```
## Show Source Files
```{r}
#| label: source-files
show_source_files(d)
```
## Show CSV Tables and Fields to Ingest
```{r}
#| label: tbls_in
d$d_csv$tables |>
datatable(caption = "Tables to ingest.")
```
```{r}
#| label: flds_in
d$d_csv$fields |>
datatable(caption = "Fields to ingest.")
```
## Show tables and fields redefined
```{r}
#| label: tbls_rd
show_tables_redefine(d)
```
```{r}
#| label: flds_rd
show_fields_redefine(d)
```
## Load Tables into Database
```{r}
#| label: load_tbls_to_db
# use ingest_dataset() which handles:
# - transform_data() for applying redefinitions
# - provenance tracking via gcs_path from read_csv_files()
# - ingest_to_working() for each table
tbl_stats <- ingest_dataset(
con = con,
d = d,
mode = if (overwrite) "replace" else "append",
verbose = TRUE
)
tbl_stats |>
datatable(rownames = FALSE, filter = "top")
```
## Apply Ship Corrections
Apply known ship name/code corrections from `ship_renames.csv`.
```{r}
#| label: apply_ship_corrections
ship_renames_csv <- here("metadata/ship_renames.csv")
d_ship_renames <- read_csv(ship_renames_csv)
# apply each correction
n_updated_total <- 0
for (i in seq_len(nrow(d_ship_renames))) {
old_name <- d_ship_renames$csv_name[i]
old_code <- d_ship_renames$csv_code[i]
new_name <- d_ship_renames$csv_name_new[i]
new_code <- d_ship_renames$csv_code_new[i]
# update where both name and code match old values
n_updated <- dbExecute(
con,
glue_sql(
"UPDATE casts
SET ship_name = {new_name}, ship_code = {new_code}
WHERE ship_name = {old_name} AND ship_code = {old_code}",
.con = con
)
)
if (n_updated > 0) {
message(glue(
"Updated {n_updated} casts: ",
"'{old_name}' [{old_code}] -> '{new_name}' [{new_code}]"
))
}
n_updated_total <- n_updated_total + n_updated
}
message(glue("\nTotal casts updated: {n_updated_total}"))
```
## Column Naming Rationale
The column renames in `flds_redefine.csv` follow these db conventions:
- **`*_id`** for integer primary/foreign keys (e.g. `Cst_Cnt` → `cast_id`, `Btl_Cnt` → `bottle_id`)
- **`*_key`** for varchar natural/composite keys (e.g. `Cast_ID` → `cast_key`, `Sta_ID` → `site_key`)
- Note: `Cruise_ID` → `cruise_key_0` (interim `_0` suffix avoids collision with SWFSC `cruise_key` YYYY-MM-NODC; discarded after merge)
- **unit suffixes** for measurements (e.g. `Depthm` → `depth_m`)
### Ambiguous column pairs
| Column pair | Why different |
|------------|---------------|
| `Cst_Cnt` (→ `cast_id`) vs `Cast_ID` (→ `cast_key`) | integer counter (1, 2, 3...) vs varchar composite (19-4903CR-HY-060-0930-05400560) |
| `Sta_ID` (→ `site_key`) vs `DbSta_ID` (→ `db_sta_key`) | "054.0 056.0" (space-separated) vs "05400560" (compact, derivable) |
| `Depth_ID` (→ `depth_key`) vs `Depthm` (→ `depth_m`) | varchar identifier vs numeric depth in meters |
### `R_*` = "Reported" (pre-QC) values
The `R_*` prefix means original instrument values before quality control.
**Critical**: `R_Sal` (Specific Volume Anomaly, values ~233, units 10⁻⁸ m³/kg)
is a **completely different parameter/scale** than `Salnty` (Practical Salinity
PSS-78, values ~33.44). These are pivoted as distinct measurement types
(`r_salinity_sva` vs `salinity`).
## Consolidate Casts Columns
Combine `date` + `time` into `datetime_utc`, drop 16 derivable columns,
and create a `casts_derived` convenience VIEW.
```{r}
#| label: consolidate_casts
# -- create datetime_utc from date + time (idempotent) ----
casts_cols <- dbListFields(con, "casts")
if ("date" %in% casts_cols && "time" %in% casts_cols) {
# date is DATE type (e.g. 1949-03-01), time is INTERVAL (e.g. 09:30:00)
# adding DATE + INTERVAL yields TIMESTAMP; 324 casts have NULL time
dbExecute(con, "ALTER TABLE casts ADD COLUMN IF NOT EXISTS datetime_utc TIMESTAMP")
dbExecute(
con,
"
UPDATE casts SET datetime_utc =
CASE
WHEN \"time\" IS NOT NULL
THEN CAST(\"date\" AS TIMESTAMP) + \"time\"
ELSE CAST(\"date\" AS TIMESTAMP)
END"
)
message(glue(
"Created datetime_utc: {dbGetQuery(con, 'SELECT COUNT(*) FROM casts WHERE datetime_utc IS NOT NULL')[[1]]} non-null values"
))
# -- drop derivable columns ----
cols_to_drop <- c(
"year", "month", "quarter", "julian_day", "julian_date",
"time_zone", "lat_deg", "lat_min", "lat_hem",
"lon_deg", "lon_min", "lon_hem",
"cruise", "cruz_sta", "db_sta_key", "cruz_num",
"date", "time"
)
n_before <- length(casts_cols)
for (col in cols_to_drop) {
tryCatch(
dbExecute(con, glue('ALTER TABLE casts DROP COLUMN "{col}"')),
error = function(e) NULL)
}
n_after <- length(dbListFields(con, "casts"))
message(glue("casts columns: {n_before} -> {n_after} (dropped {n_before - n_after})"))
} else {
message("casts already consolidated (datetime_utc exists, date/time dropped)")
}
```
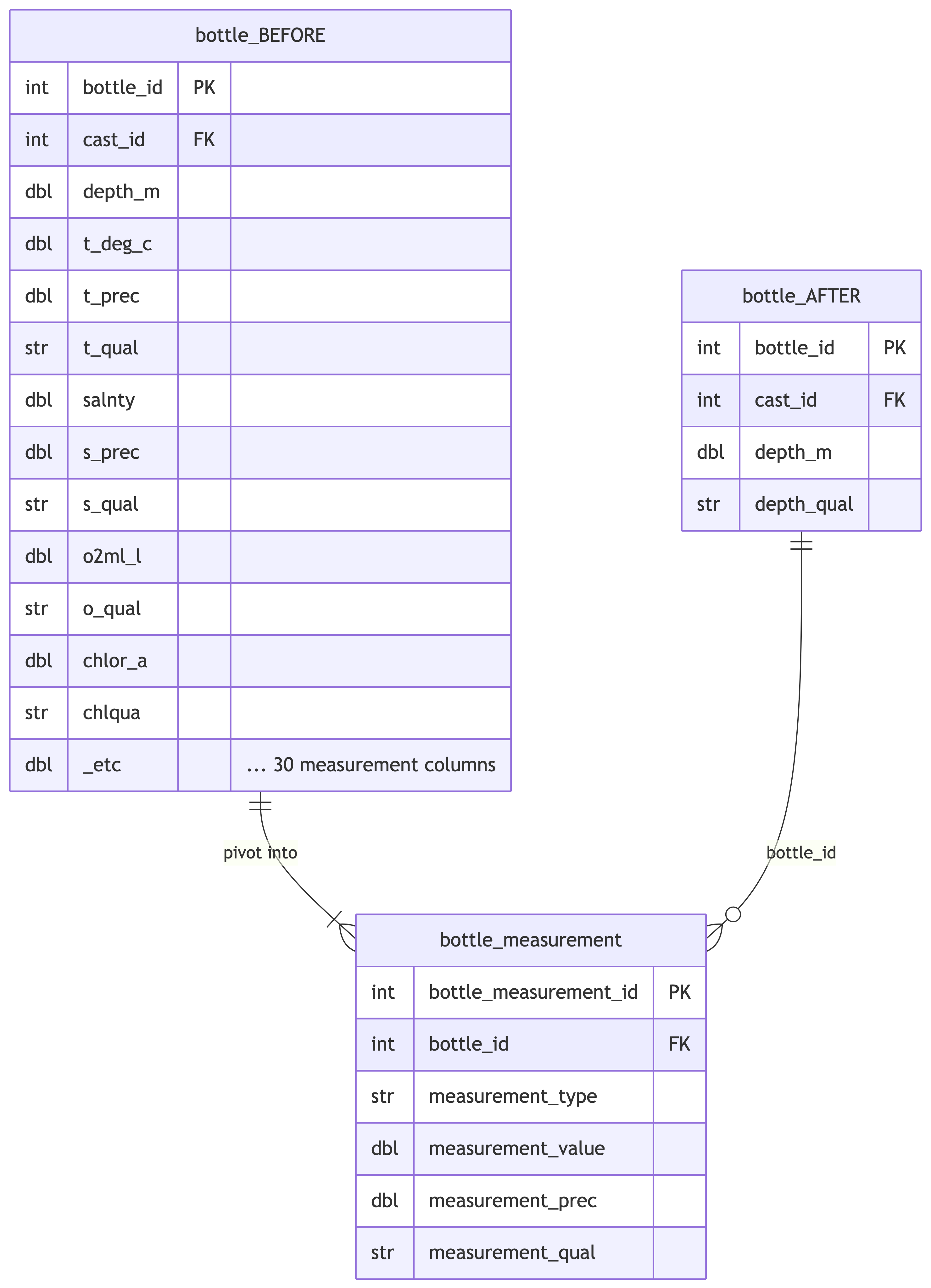
## Pivot Bottle Measurements
Pivot wide-format measurement columns from `bottle` into long-format
`bottle_measurement` table. Each measurement type has a value, optional
precision, and optional quality code.
```{mermaid}
%%| label: pivot_bottle_meas_diagram
%%| fig-cap: "Pivot wide bottle columns into long-format bottle_measurement table."
erDiagram
bottle_BEFORE {
int bottle_id PK
int cast_id FK
dbl depth_m
dbl t_deg_c
dbl t_prec
str t_qual
dbl salnty
dbl s_prec
str s_qual
dbl o2ml_l
str o_qual
dbl chlor_a
str chlqua
dbl _etc "... 30 measurement columns"
}
bottle_AFTER {
int bottle_id PK
int cast_id FK
dbl depth_m
str depth_qual
}
bottle_measurement {
int bottle_measurement_id PK
int bottle_id FK
str measurement_type
dbl measurement_value
dbl measurement_prec
str measurement_qual
}
bottle_BEFORE ||--|{ bottle_measurement : "pivot into"
bottle_AFTER ||--o{ bottle_measurement : "bottle_id"
```
```{r}
#| label: pivot_bottle_measurements
# build measurement mapping from unified CSV
bottle_meas_map <- d_meas_type |>
filter(
str_detect(`_source_datasets`, "calcofi_bottle"),
`_source_table` == "bottle"
) |>
select(
measurement_type,
value_col = `_source_column`,
prec_col = `_prec_column`,
qual_col = `_qual_column`
) |>
mutate(across(c(prec_col, qual_col), ~ na_if(.x, "")))
# build UNION ALL SQL from mapping
sql_parts <- purrr::pmap_chr(
bottle_meas_map,
function(
measurement_type,
value_col,
prec_col,
qual_col
) {
prec_expr <- if (is.na(prec_col)) "NULL" else prec_col
qual_expr <- if (is.na(qual_col)) "NULL" else qual_col
glue(
"SELECT bottle_id, '{measurement_type}' AS measurement_type,
CAST({value_col} AS DOUBLE) AS measurement_value,
CAST({prec_expr} AS DOUBLE) AS measurement_prec,
CAST({qual_expr} AS VARCHAR) AS measurement_qual
FROM bottle WHERE {value_col} IS NOT NULL"
)
}
)
sql_create <- glue(
"CREATE OR REPLACE TABLE bottle_measurement AS
SELECT ROW_NUMBER() OVER (ORDER BY bottle_id, measurement_type)
AS bottle_measurement_id, *
FROM (
{paste(sql_parts, collapse = '\nUNION ALL\n')}
) sub"
)
dbExecute(con, sql_create)
n_meas <- dbGetQuery(con, "SELECT COUNT(*) FROM bottle_measurement")[[1]]
n_types <- dbGetQuery(
con,
"SELECT COUNT(DISTINCT measurement_type) FROM bottle_measurement"
)[[1]]
message(glue(
"bottle_measurement: {format(n_meas, big.mark = ',')} rows, {n_types} measurement types"
))
# -- drop measurement columns from bottle base table ----
cols_to_drop <- unique(c(
bottle_meas_map$value_col,
na.omit(bottle_meas_map$prec_col),
na.omit(bottle_meas_map$qual_col)
))
n_before <- length(dbListFields(con, "bottle"))
for (col in cols_to_drop) {
tryCatch(
dbExecute(con, glue('ALTER TABLE bottle DROP COLUMN "{col}"')),
error = function(e) warning(glue("could not drop {col}: {e$message}"))
)
}
# rename p_qual to depth_qual (pressure/depth quality code stays in base table)
dbExecute(con, 'ALTER TABLE bottle RENAME COLUMN p_qual TO depth_qual')
n_after <- length(dbListFields(con, "bottle"))
message(glue(
"bottle columns: {n_before} -> {n_after} (dropped {n_before - n_after})"
))
```
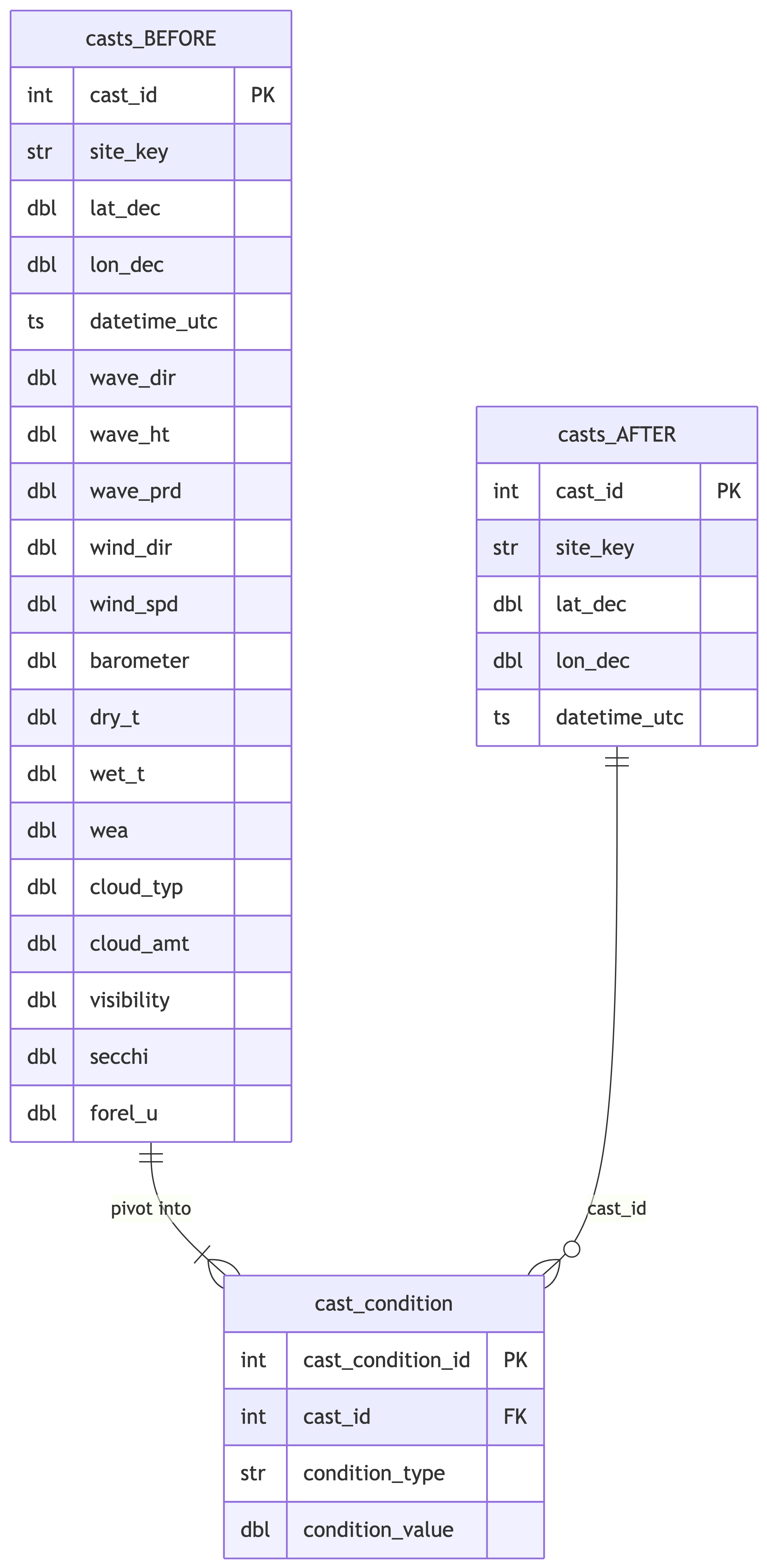
## Pivot Cast Conditions
Pivot meteorological and environmental observation columns from `casts`
into long-format `cast_condition` table.
```{mermaid}
%%| label: pivot_cast_cond_diagram
%%| fig-cap: "Pivot wide cast condition columns into long-format cast_condition table."
erDiagram
casts_BEFORE {
int cast_id PK
str site_key
dbl lat_dec
dbl lon_dec
ts datetime_utc
dbl wave_dir
dbl wave_ht
dbl wave_prd
dbl wind_dir
dbl wind_spd
dbl barometer
dbl dry_t
dbl wet_t
dbl wea
dbl cloud_typ
dbl cloud_amt
dbl visibility
dbl secchi
dbl forel_u
}
casts_AFTER {
int cast_id PK
str site_key
dbl lat_dec
dbl lon_dec
ts datetime_utc
}
cast_condition {
int cast_condition_id PK
int cast_id FK
str condition_type
dbl condition_value
}
casts_BEFORE ||--|{ cast_condition : "pivot into"
casts_AFTER ||--o{ cast_condition : "cast_id"
```
```{r}
#| label: pivot_cast_conditions
# build cast condition mapping from unified CSV
cast_cond_map <- d_meas_type |>
filter(
str_detect(`_source_datasets`, "calcofi_bottle"),
`_source_table` == "casts"
) |>
select(
condition_type = measurement_type,
source_col = `_source_column`
)
# build UNION ALL SQL
sql_parts <- purrr::pmap_chr(
cast_cond_map,
function(condition_type, source_col) {
glue(
"SELECT cast_id, '{condition_type}' AS condition_type,
CAST({source_col} AS DOUBLE) AS condition_value
FROM casts WHERE {source_col} IS NOT NULL"
)
}
)
sql_create <- glue(
"CREATE OR REPLACE TABLE cast_condition AS
SELECT ROW_NUMBER() OVER (ORDER BY cast_id, condition_type)
AS cast_condition_id, *
FROM (
{paste(sql_parts, collapse = '\nUNION ALL\n')}
) sub"
)
dbExecute(con, sql_create)
n_cond <- dbGetQuery(con, "SELECT COUNT(*) FROM cast_condition")[[1]]
n_types <- dbGetQuery(
con,
"SELECT COUNT(DISTINCT condition_type) FROM cast_condition"
)[[1]]
message(glue(
"cast_condition: {format(n_cond, big.mark = ',')} rows, {n_types} condition types"
))
# -- drop condition columns from casts ----
n_before <- length(dbListFields(con, "casts"))
for (col in cast_cond_map$source_col) {
tryCatch(
dbExecute(con, glue('ALTER TABLE casts DROP COLUMN "{col}"')),
error = function(e) warning(glue("could not drop {col}: {e$message}"))
)
}
n_after <- length(dbListFields(con, "casts"))
message(glue(
"casts columns: {n_before} -> {n_after} (dropped {n_before - n_after})"
))
```
## Load Measurement Type Reference
Load the `measurement_type` lookup table that provides units and
descriptions for all measurement/condition types across datasets.
### `measurement_type` vs `lookup`
The `measurement_type` table and the swfsc `lookup` table serve fundamentally different purposes:
- **`measurement_type`** answers "what was measured?" — provides metadata (units, description, source provenance) for measurement codes used in `bottle_measurement`, `cast_condition`, and `ichthyo`
- **`lookup`** answers "what does this code mean?" — translates categorical values (egg developmental stages 1-11, larva stages, tow types) into human-readable descriptions
Merging would create a sparse table with many NULL columns (e.g., `units`/`_source_column` NULL for egg stages; `lookup_num`/`lookup_chr` NULL for temperature). Both tables coexist in the Working DuckLake, loaded from their respective upstream workflows.
```{r}
#| label: load_measurement_type
dbWriteTable(con, "measurement_type", d_meas_type, overwrite = TRUE)
# validate FK: all bottle_measurement types exist in reference table
orphan_btl_types <- dbGetQuery(
con,
"
SELECT DISTINCT measurement_type FROM bottle_measurement
WHERE measurement_type NOT IN (SELECT measurement_type FROM measurement_type)"
)
stopifnot(
"all bottle_measurement.measurement_type must exist in measurement_type" = nrow(
orphan_btl_types
) ==
0
)
# validate FK: all cast_condition types exist in reference table
orphan_cond_types <- dbGetQuery(
con,
"
SELECT DISTINCT condition_type FROM cast_condition
WHERE condition_type NOT IN (SELECT measurement_type FROM measurement_type)"
)
stopifnot(
"all cast_condition.condition_type must exist in measurement_type" = nrow(
orphan_cond_types
) ==
0
)
message(glue(
"measurement_type: {nrow(d_meas_type)} types loaded, FK checks passed"
))
# validate: all measurement_types used by this dataset are registered
bottle_types_used <- dbGetQuery(
con,
"SELECT DISTINCT measurement_type FROM bottle_measurement"
)$measurement_type
cast_types_used <- dbGetQuery(
con,
"SELECT DISTINCT condition_type FROM cast_condition"
)$condition_type
registered <- d_meas_type |>
filter(str_detect(`_source_datasets`, "calcofi_bottle")) |>
pull(measurement_type)
unregistered <- setdiff(c(bottle_types_used, cast_types_used), registered)
stopifnot(
"all measurement_types used by calcofi_bottle must be registered in _source_datasets" = length(
unregistered
) ==
0
)
```
## Load Dataset Metadata
```{r}
#| label: load-dataset-metadata
d_dataset <- read_csv(here("metadata/dataset.csv"))
dbWriteTable(con, "dataset", d_dataset, overwrite = TRUE)
message(glue("dataset: {nrow(d_dataset)} datasets registered"))
```
## Verify Primary Keys
```{r}
#| label: verify_primary_keys
# verify cast_id uniqueness in casts
cast_keys <- tbl(con, "casts") |> pull(cast_id)
n_dup_cast <- sum(duplicated(cast_keys))
stopifnot("cast_id must be unique in casts" = n_dup_cast == 0)
message(glue("casts.cast_id: {length(cast_keys)} unique values, 0 duplicates"))
# verify bottle_id uniqueness in bottle
btl_keys <- tbl(con, "bottle") |> pull(bottle_id)
n_dup_btl <- sum(duplicated(btl_keys))
stopifnot("bottle_id must be unique in bottle" = n_dup_btl == 0)
message(glue(
"bottle.bottle_id: {length(btl_keys)} unique values, 0 duplicates"
))
# verify FK: bottle.cast_id -> casts.cast_id
orphan_bottles <- tbl(con, "bottle") |>
anti_join(
tbl(con, "casts"),
by = "cast_id"
) |>
collect()
stopifnot(
"all bottle.cast_id must exist in casts.cast_id" = nrow(orphan_bottles) == 0
)
message(glue(
"FK integrity: all {length(btl_keys)} bottle.cast_id ",
"found in casts.cast_id"
))
# verify bottle_measurement_id uniqueness
n_dup_bm_id <- dbGetQuery(
con,
"
SELECT COUNT(*) FROM (
SELECT bottle_measurement_id, COUNT(*) AS n
FROM bottle_measurement
GROUP BY bottle_measurement_id
HAVING COUNT(*) > 1)"
)[[1]]
stopifnot("bottle_measurement_id must be unique" = n_dup_bm_id == 0)
message(glue("bottle_measurement.bottle_measurement_id: unique, 0 duplicates"))
# verify composite PK: bottle_measurement (bottle_id, measurement_type)
n_dup_meas <- dbGetQuery(
con,
"
SELECT COUNT(*) FROM (
SELECT bottle_id, measurement_type, COUNT(*) AS n
FROM bottle_measurement
GROUP BY bottle_id, measurement_type
HAVING COUNT(*) > 1)"
)[[1]]
stopifnot(
"bottle_measurement (bottle_id, measurement_type) must be unique" = n_dup_meas ==
0
)
message(glue("bottle_measurement: composite key unique, 0 duplicates"))
# verify FK: bottle_measurement.bottle_id -> bottle.bottle_id
n_orphan_meas <- dbGetQuery(
con,
"
SELECT COUNT(DISTINCT bottle_id) FROM bottle_measurement
WHERE bottle_id NOT IN (SELECT bottle_id FROM bottle)"
)[[1]]
stopifnot(
"all bottle_measurement.bottle_id must exist in bottle" = n_orphan_meas == 0
)
message(glue("bottle_measurement FK: all bottle_ids found in bottle"))
# verify cast_condition_id uniqueness
n_dup_cc_id <- dbGetQuery(
con,
"
SELECT COUNT(*) FROM (
SELECT cast_condition_id, COUNT(*) AS n
FROM cast_condition
GROUP BY cast_condition_id
HAVING COUNT(*) > 1)"
)[[1]]
stopifnot("cast_condition_id must be unique" = n_dup_cc_id == 0)
message(glue("cast_condition.cast_condition_id: unique, 0 duplicates"))
# verify composite PK: cast_condition (cast_id, condition_type)
n_dup_cond <- dbGetQuery(
con,
"
SELECT COUNT(*) FROM (
SELECT cast_id, condition_type, COUNT(*) AS n
FROM cast_condition
GROUP BY cast_id, condition_type
HAVING COUNT(*) > 1)"
)[[1]]
stopifnot(
"cast_condition (cast_id, condition_type) must be unique" = n_dup_cond == 0
)
message(glue("cast_condition: composite key unique, 0 duplicates"))
# verify FK: cast_condition.cast_id -> casts.cast_id
n_orphan_cond <- dbGetQuery(
con,
"
SELECT COUNT(DISTINCT cast_id) FROM cast_condition
WHERE cast_id NOT IN (SELECT cast_id FROM casts)"
)[[1]]
stopifnot("all cast_condition.cast_id must exist in casts" = n_orphan_cond == 0)
message(glue("cast_condition FK: all cast_ids found in casts"))
```
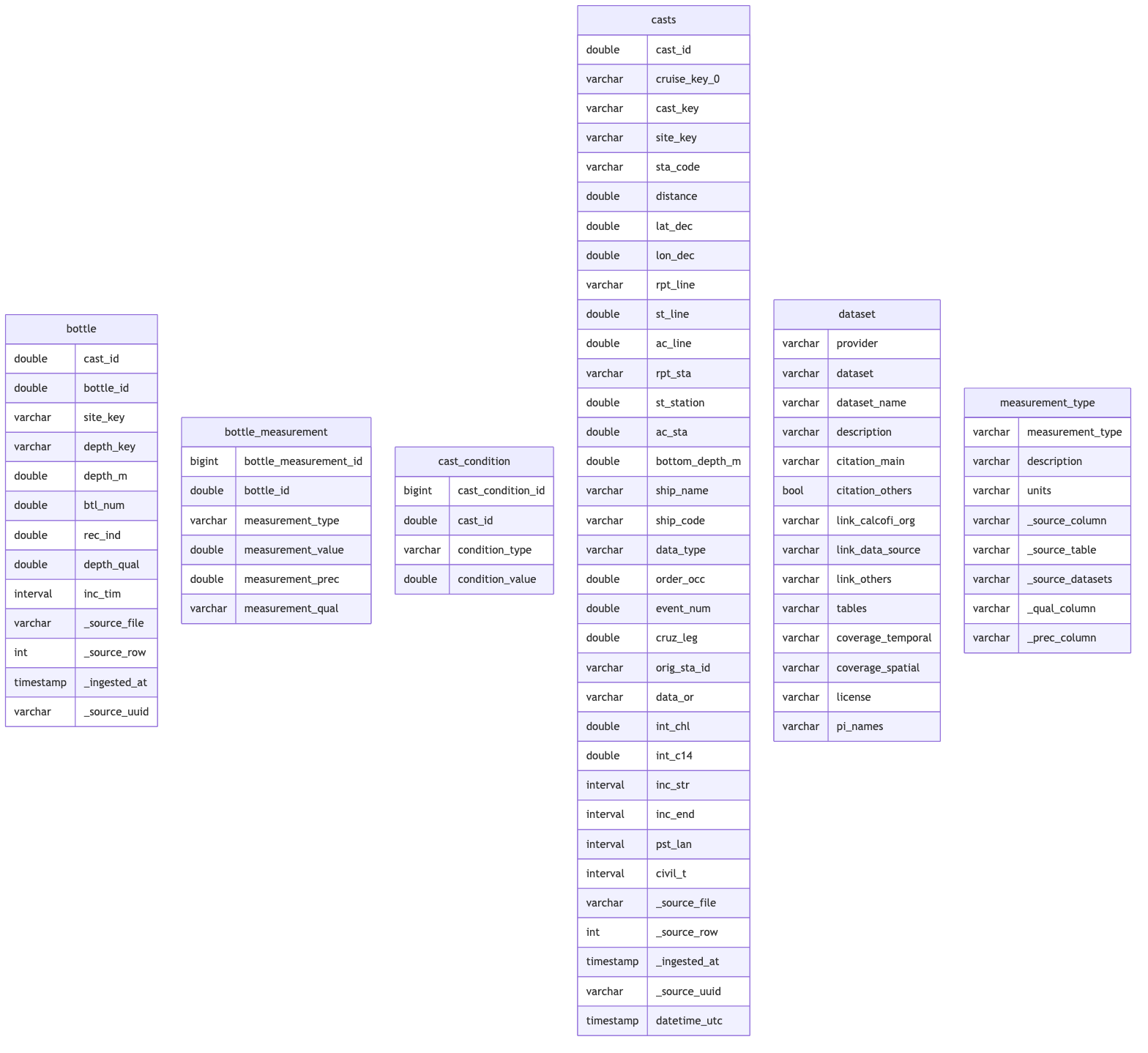
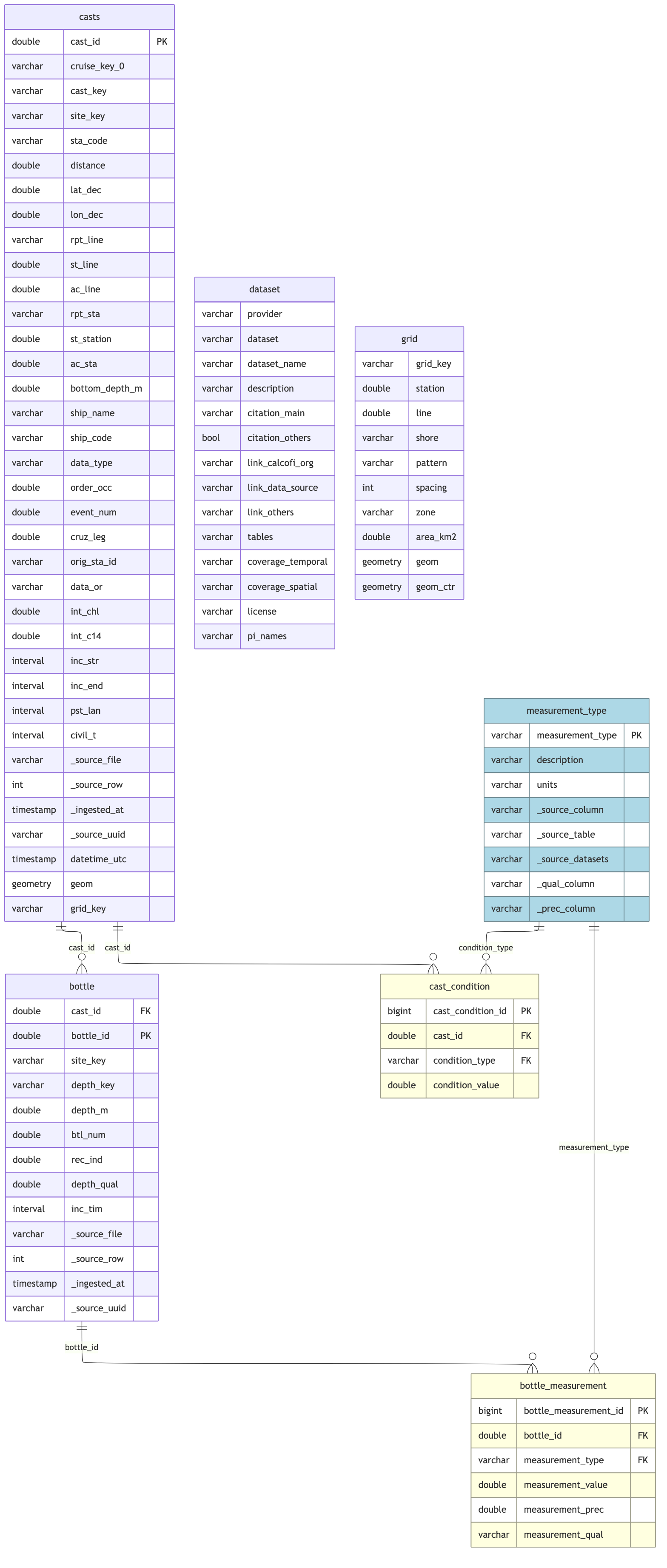
## Schema Documentation
```{r}
#| label: erd_tbls
# exclude views (casts_derived)
db_tables <- function(con) {
setdiff(dbListTables(con), "casts_derived") |> sort()
}
tbls <- db_tables(con)
cc_erd(con, tables = tbls)
```
### Primary Key Strategy
**Note on UUID-first approach**: The SWFSC ichthyo tables use source UUIDs (minted
at sea) as primary identifiers. The bottle/cast tables are different — they use
source integer counters (`Cst_Cnt`, `Btl_Cnt`) as natural keys, which are stable
identifiers from the source system. The pivoted tables (`bottle_measurement`,
`cast_condition`) have no source identifier and use sequential integer IDs for
convenience, with composite natural keys enforced as unique.
| Table | Primary Key | Type |
|-------|-------------|------|
| `casts` | `cast_id` | Source integer counter (`Cst_Cnt`) — stable source key |
| `bottle` | `bottle_id` | Source integer counter (`Btl_Cnt`) — stable source key |
| `bottle_measurement` | `bottle_measurement_id` | Sequential (derived/pivoted table); composite `(bottle_id, measurement_type)` enforced unique |
| `cast_condition` | `cast_condition_id` | Sequential (derived/pivoted table); composite `(cast_id, condition_type)` enforced unique |
| `measurement_type` | `measurement_type` | Natural key (type code string) |
| `grid` | `grid_key` | Natural key |
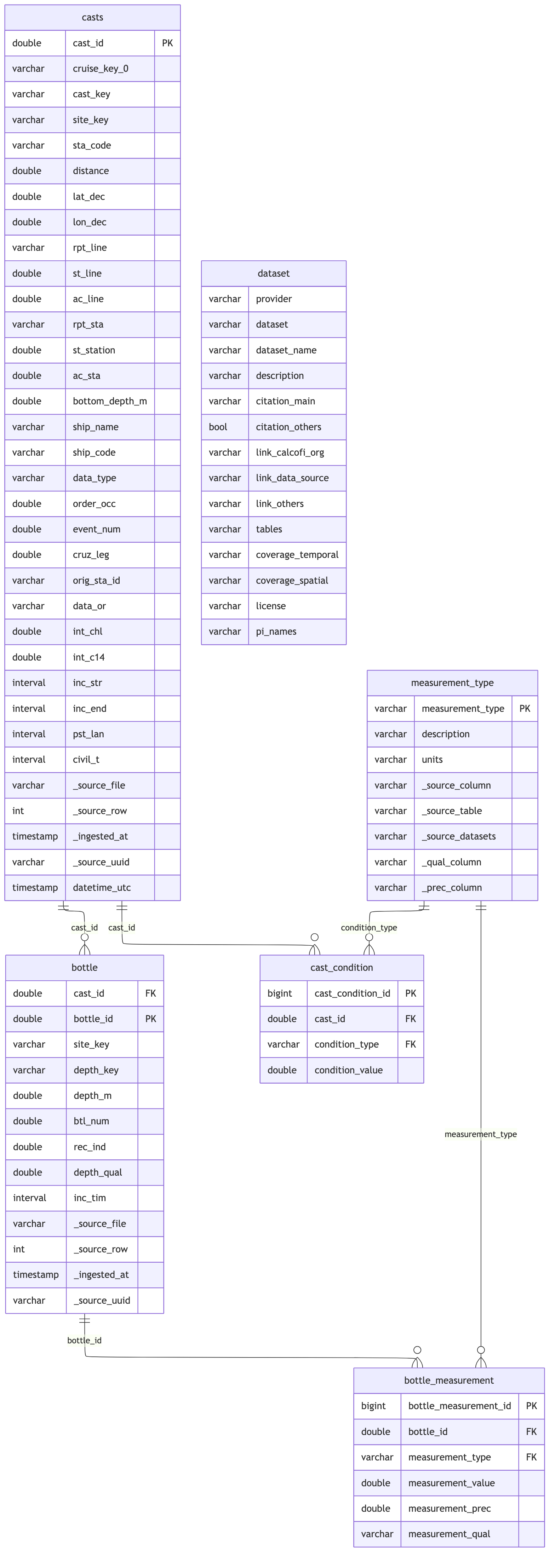
### Foreign Key Relationships
```
casts.cast_id (PK, integer) — station-level
↓
bottle.bottle_id (PK, integer) — sample-level
bottle.cast_id (FK) → casts.cast_id
↓
bottle_measurement.bottle_measurement_id (PK, integer) — ROW_NUMBER(ORDER BY bottle_id, measurement_type)
bottle_measurement.bottle_id (FK) → bottle.bottle_id
bottle_measurement.measurement_type (FK) → measurement_type.measurement_type
cast_condition.cast_condition_id (PK, integer) — ROW_NUMBER(ORDER BY cast_id, condition_type)
cast_condition.cast_id (FK) → casts.cast_id
cast_condition.condition_type (FK) → measurement_type.measurement_type
casts.grid_key (FK) → grid.grid_key (after spatial join)
casts.ship_code (soft FK) → ship.ship_nodc (validated after integration with swfsc)
```
```{r}
#| label: erd_fk
# define PK/FK relationships for visualization and relationships.json
bottle_rels <- list(
primary_keys = list(
casts = "cast_id",
bottle = "bottle_id",
bottle_measurement = "bottle_measurement_id",
cast_condition = "cast_condition_id",
measurement_type = "measurement_type"
),
foreign_keys = list(
list(
table = "bottle",
column = "cast_id",
ref_table = "casts",
ref_column = "cast_id"
),
list(
table = "bottle_measurement",
column = "bottle_id",
ref_table = "bottle",
ref_column = "bottle_id"
),
list(
table = "bottle_measurement",
column = "measurement_type",
ref_table = "measurement_type",
ref_column = "measurement_type"
),
list(
table = "cast_condition",
column = "cast_id",
ref_table = "casts",
ref_column = "cast_id"
),
list(
table = "cast_condition",
column = "condition_type",
ref_table = "measurement_type",
ref_column = "measurement_type"
)
)
)
cc_erd(con, tables = db_tables(con), rels = bottle_rels)
```
## Add Spatial
### Add `casts.geom`
```{r}
#| label: mk_cast_pts
# add geometry column using DuckDB spatial
# note: DuckDB spatial doesn't track SRID metadata (unlike PostGIS)
# all geometries assumed WGS84 (EPSG:4326) by convention
add_point_geom(con, "casts")
```
### Load `grid` table
Load grid from the ichthyo workflow's local parquet output (single source of
truth — grid is created in `ingest_swfsc_ichthyo.qmd`).
```{r}
#| label: load_grid
load_prior_tables(
con = con,
tables = "grid",
parquet_dir = here("data/parquet/swfsc_ichthyo"),
as_view = TRUE
)
```
### Update `casts.grid_key`
```{r}
#| label: update_cast_from_grid
grid_stats <- assign_grid_key(con, "casts")
grid_stats |> datatable()
```
### Create `casts_derived` VIEW
Create convenience VIEW after all casts table modifications are complete
(column drops, pivots, spatial join). The VIEW adds back temporal and
coordinate columns derivable from the base table.
```{r}
#| label: create_casts_derived
dbExecute(con, "DROP VIEW IF EXISTS casts_derived")
dbExecute(
con,
"
CREATE VIEW casts_derived AS
SELECT *,
EXTRACT(YEAR FROM datetime_utc)::SMALLINT AS year,
EXTRACT(MONTH FROM datetime_utc)::SMALLINT AS month,
EXTRACT(QUARTER FROM datetime_utc)::SMALLINT AS quarter,
EXTRACT(DOY FROM datetime_utc)::SMALLINT AS julian_day,
(datetime_utc::DATE - DATE '1899-12-30') AS julian_date,
FLOOR(ABS(lat_dec))::SMALLINT AS lat_deg,
(ABS(lat_dec) - FLOOR(ABS(lat_dec))) * 60 AS lat_min,
CASE WHEN lat_dec >= 0 THEN 'N' ELSE 'S' END AS lat_hem,
FLOOR(ABS(lon_dec))::SMALLINT AS lon_deg,
(ABS(lon_dec) - FLOOR(ABS(lon_dec))) * 60 AS lon_min,
CASE WHEN lon_dec >= 0 THEN 'E' ELSE 'W' END AS lon_hem,
STRFTIME(datetime_utc, '%Y%m') AS cruise,
REPLACE(REPLACE(site_key, '.', ''), ' ', '') AS db_sta_key
FROM casts"
)
message("Created casts_derived VIEW with derived temporal/coordinate columns")
```
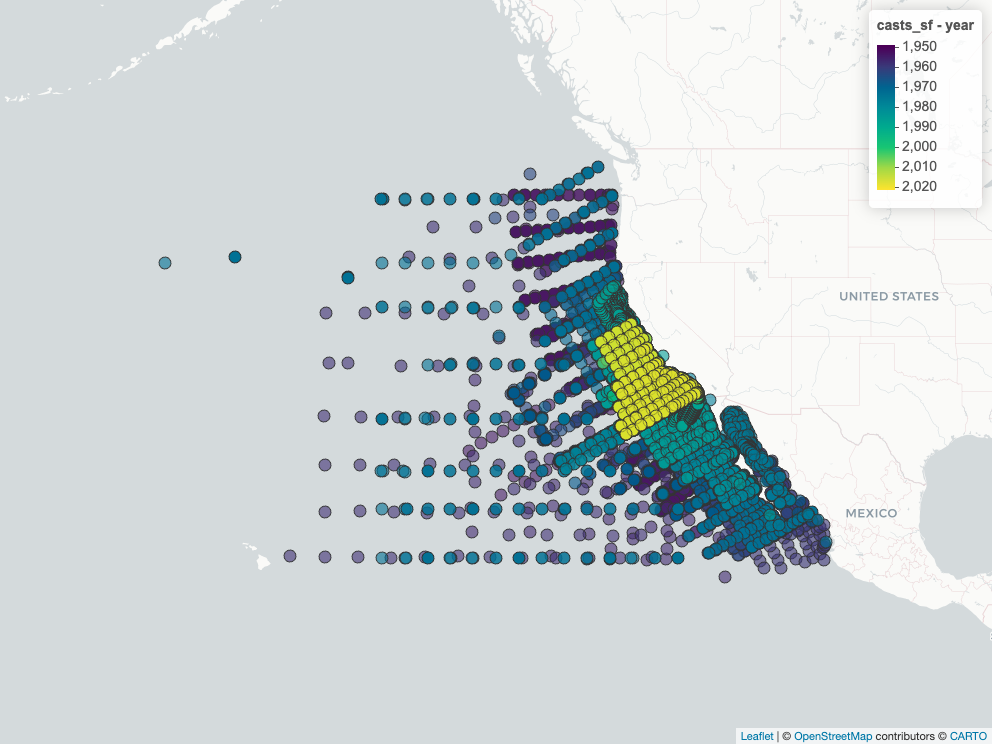
### Map Cast Locations
```{r}
#| label: map_casts
# slowish, so use cached figure
map_casts_png <- here(glue("figures/{provider}_{dataset}_casts_map.png"))
if (!file_exists(map_casts_png)) {
# exclude native GEOMETRY column; convert via ST_AsText for sf
cast_cols <- dbGetQuery(
con,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'casts' AND data_type != 'GEOMETRY'"
)$column_name
cast_sql <- paste(
"SELECT",
paste(cast_cols, collapse = ", "),
", ST_AsText(geom) AS geom_wkt FROM casts"
)
casts_sf <- dbGetQuery(con, cast_sql) |>
st_as_sf(wkt = "geom_wkt", crs = 4326) |>
select(-geom_wkt) |>
mutate(year = year(datetime_utc))
m <- mapView(casts_sf, zcol = "year")
mapshot2(m, file = map_casts_png)
}
htmltools::img(
src = map_casts_png |> str_replace(here(), "."),
width = "600px"
)
```
## Report
```{r}
#| label: show_latest
# cc_erd handles GEOMETRY columns natively (unlike dm_from_con)
cc_erd(
con,
tables = db_tables(con),
rels = bottle_rels,
colors = list(
lightyellow = c("bottle_measurement", "cast_condition"),
lightblue = "measurement_type"
)
)
```
```{r}
#| label: summary_stats
# summary statistics
n_casts <- dbGetQuery(con, "SELECT COUNT(*) FROM casts")[[1]]
n_bottles <- tbl(con, "bottle") |> tally() |> pull(n)
n_meas <- dbGetQuery(con, "SELECT COUNT(*) FROM bottle_measurement")[[1]]
n_cond <- dbGetQuery(con, "SELECT COUNT(*) FROM cast_condition")[[1]]
year_range <- dbGetQuery(
con,
"
SELECT
EXTRACT(YEAR FROM MIN(datetime_utc))::INTEGER AS yr_min,
EXTRACT(YEAR FROM MAX(datetime_utc))::INTEGER AS yr_max
FROM casts"
)
fmt <- function(x, ...) format(x, big.mark = ",", ...)
message(glue("Total casts: {fmt(n_casts)}"))
message(glue("Total bottles: {fmt(n_bottles)}"))
message(glue("Total bottle measurements: {fmt(n_meas)}"))
message(glue("Total cast conditions: {fmt(n_cond)}"))
message(glue("Year range: {year_range$yr_min} - {year_range$yr_max}"))
```
## Validate Local Database
Validate data quality in the local wrangling database before exporting
to parquet. The parquet outputs from this workflow can later be used to
update the Working DuckLake.
```{r}
#| label: validate
# validate data quality
validation <- validate_for_release(con)
if (validation$passed) {
message("Validation passed!")
if (nrow(validation$checks) > 0) {
validation$checks |>
filter(status != "pass") |>
datatable(caption = "Validation Warnings")
}
} else {
cat("Validation FAILED:\n")
cat(paste("-", validation$errors, collapse = "\n"))
}
```
## Enforce Column Types
Force integer/smallint types on columns that R's `numeric` mapped to
`DOUBLE` during `dbWriteTable()`. Uses `flds_redefine.csv` (`type_new`)
as the source of truth for source-table columns, plus explicit overrides
for derived-table columns.
```{r}
#| label: enforce_types
type_changes <- enforce_column_types(
con = con,
d_flds_rd = d$d_flds_rd,
type_overrides = list(),
tables = db_tables(con),
verbose = TRUE
)
if (nrow(type_changes) > 0) {
type_changes |>
datatable(caption = "Column type changes applied")
}
# recreate casts_derived VIEW after type changes
dbExecute(con, "DROP VIEW IF EXISTS casts_derived")
dbExecute(
con,
"
CREATE VIEW casts_derived AS
SELECT *,
EXTRACT(YEAR FROM datetime_utc)::SMALLINT AS year,
EXTRACT(MONTH FROM datetime_utc)::SMALLINT AS month,
EXTRACT(QUARTER FROM datetime_utc)::SMALLINT AS quarter,
EXTRACT(DOY FROM datetime_utc)::SMALLINT AS julian_day,
(datetime_utc::DATE - DATE '1899-12-30') AS julian_date,
FLOOR(ABS(lat_dec))::SMALLINT AS lat_deg,
(ABS(lat_dec) - FLOOR(ABS(lat_dec))) * 60 AS lat_min,
CASE WHEN lat_dec >= 0 THEN 'N' ELSE 'S' END AS lat_hem,
FLOOR(ABS(lon_dec))::SMALLINT AS lon_deg,
(ABS(lon_dec) - FLOOR(ABS(lon_dec))) * 60 AS lon_min,
CASE WHEN lon_dec >= 0 THEN 'E' ELSE 'W' END AS lon_hem,
STRFTIME(datetime_utc, '%Y%m') AS cruise,
REPLACE(REPLACE(site_key, '.', ''), ' ', '') AS db_sta_key
FROM casts"
)
message("Recreated casts_derived VIEW after column type enforcement")
```
## Data Preview
Preview first and last rows of each table before writing parquet outputs.
```{r}
#| label: preview_tables
#| results: asis
preview_tables(
con,
c(
"casts",
"bottle",
"bottle_measurement",
"cast_condition",
"measurement_type"
)
) # grid canonical from swfsc
```
```{r}
#| label: preview_casts_derived
# casts_derived is a VIEW; exclude GEOMETRY column for R driver compatibility
cd_cols <- dbGetQuery(
con,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'casts_derived' AND data_type != 'GEOMETRY'"
)$column_name
dbGetQuery(
con,
glue("SELECT {paste(cd_cols, collapse=', ')} FROM casts_derived LIMIT 100")
) |>
datatable(
caption = "casts_derived — first 100 rows (VIEW)",
rownames = FALSE,
filter = "top"
)
```
## Write Parquet Outputs
Export tables to parquet files for downstream use.
```{r}
#| label: write_parquet
# collect mismatches for manifest
mismatches <- list(
ships = collect_ship_mismatches(con, "casts"),
measurement_types = collect_measurement_type_mismatches(
con,
here("metadata/measurement_type.csv")
),
cruise_keys = collect_cruise_key_mismatches(con, "casts")
)
# write parquet files with manifest
# export only tables this ingest creates (dependency _new deltas added later)
own_tables <- c(
"casts",
"bottle",
"bottle_measurement",
"cast_condition",
"measurement_type",
"dataset"
)
parquet_stats <- write_parquet_outputs(
con = con,
output_dir = dir_parquet,
tables = own_tables,
sort_by = list(
bottle_measurement = "measurement_type",
casts = "hilbert:lon_dec,lat_dec"),
strip_provenance = FALSE,
mismatches = mismatches
)
parquet_stats |>
mutate(file = basename(path)) |>
select(-path) |>
datatable(caption = "Parquet export statistics")
```
## Write Metadata
Build `metadata.json` sidecar file documenting all tables and columns in
parquet outputs. DuckDB `COMMENT ON` does not propagate to parquet, so
this provides the metadata externally.
```{r}
#| label: write_metadata
metadata_path <- build_metadata_json(
con = con,
d_tbls_rd = d$d_tbls_rd,
d_flds_rd = d$d_flds_rd,
metadata_derived_csv = here(
"metadata/calcofi/bottle/metadata_derived.csv"
),
output_dir = dir_parquet,
tables = db_tables(con),
set_comments = TRUE,
provider = provider,
dataset = dataset,
workflow_url = glue(
"https://calcofi.io/workflows/",
"ingest_calcofi_bottle.html"
)
)
# write relationships.json sidecar with PKs/FKs
build_relationships_json(
rels = bottle_rels,
output_dir = dir_parquet,
provider = provider,
dataset = dataset
)
# show metadata summary
metadata <- jsonlite::fromJSON(metadata_path)
tibble(
table = names(metadata$tables),
n_cols = map_int(
names(metadata$tables),
~ sum(grepl(glue("^{.x}\\."), names(metadata$columns)))
),
name_long = map_chr(metadata$tables, ~ .x$name_long)
) |>
datatable(caption = "Table metadata summary")
```
```{r}
#| label: show_metadata_json
listviewer::jsonedit(
jsonlite::fromJSON(metadata_path, simplifyVector = FALSE),
mode = "view")
```
```{r}
#| label: show_relationships_json
listviewer::jsonedit(
jsonlite::fromJSON(
file.path(dir_parquet, "relationships.json"),
simplifyVector = FALSE),
mode = "view")
```
## Upload to GCS Archive
Upload parquet files, manifest, and metadata sidecar to
`gs://calcofi-db/ingest/{provider}_{dataset}/`.
```{r}
#| label: upload_gcs
sync_to_gcs(
local_dir = dir_parquet,
gcs_prefix = glue("ingest/{dir_label}"),
bucket = "calcofi-db")
```
## Cross-Dataset Integration
Link bottle casts to SWFSC ship/cruise reference tables via ship matching
and cruise key derivation. This replaces the separate `merge_ichthyo_bottle.qmd`
workflow step.
### Load Reference Tables
```{r}
#| label: load_reference_tables
# tables declared in calcofi.modifies frontmatter — loaded as TABLEs (writable)
# all other dependency tables loaded as VIEWs (zero-copy, read-only)
modifies_tables <- c("ship") # from calcofi.modifies in YAML frontmatter
load_prior_tables(
con = con,
tables = modifies_tables,
parquet_dir = here("data/parquet/swfsc_ichthyo")
)
load_prior_tables(
con = con,
tables = setdiff(c("ship", "cruise"), modifies_tables),
parquet_dir = here("data/parquet/swfsc_ichthyo"),
as_view = TRUE
)
# snapshot PKs of modified tables before modifications
modifies_pks <- list()
for (tbl in modifies_tables) {
pk_col <- dbGetQuery(
con,
glue(
"SELECT column_name FROM information_schema.columns
WHERE table_name = '{tbl}' ORDER BY ordinal_position LIMIT 1"
)
)$column_name
modifies_pks[[tbl]] <- list(
pk_col = pk_col,
keys = dbGetQuery(con, glue("SELECT {pk_col} FROM {tbl}"))[[1]]
)
}
```
### Ship Matching + Cruise Bridge
```{r}
#| label: derive_cruise_key
bridge_result <- derive_cruise_key_on_casts(
con = con,
ship_renames_csv = here("metadata/ship_renames.csv"),
fetch_ices = TRUE
)
# insert interim entries for still-unmatched ships (ship_nodc = "?XX?")
ensure_interim_ships(con, bridge_result$ship_matches)
bridge_result$ship_matches |>
datatable(caption = "Ship matching results")
```
```{r}
#| label: ship_match_stats
bridge_result$ship_matches |>
count(match_type) |>
datatable(caption = "Ship match type counts")
```
```{r}
#| label: cruise_bridge_stats
bridge_result$cruise_stats |>
datatable(caption = "Cruise bridge match statistics")
```
```{r}
#| label: unmatched_ships
if (nrow(bridge_result$unmatched_report) > 0) {
bridge_result$unmatched_report |>
datatable(caption = "Unmatched ship codes (no ship_key)")
} else {
message("All ship codes matched!")
}
```
### Cross-Dataset Validation
```{r}
#| label: cross_validate
# use information_schema to check columns (avoids GEOMETRY type issues with dbListFields)
casts_cols <- dbGetQuery(
con,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'casts'"
)$column_name
grid_cols <- dbGetQuery(
con,
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'grid'"
)$column_name
# grid_key integrity: casts.grid_key should all be in grid.grid_key
if ("grid_key" %in% casts_cols && "grid_key" %in% grid_cols) {
grid_orphans <- dbGetQuery(
con,
"SELECT COUNT(*) AS n FROM casts c
WHERE c.grid_key IS NOT NULL
AND c.grid_key NOT IN (SELECT grid_key FROM grid)"
)$n
message(glue("Grid key orphans in casts: {grid_orphans}"))
}
# ship PK uniqueness
ship_dups <- dbGetQuery(
con,
"SELECT ship_key, COUNT(*) AS n FROM ship
GROUP BY ship_key HAVING COUNT(*) > 1"
)
if (nrow(ship_dups) > 0) {
warning(glue("Duplicate ship_key values: {nrow(ship_dups)}"))
}
# cruise PK uniqueness
cruise_dups <- dbGetQuery(
con,
"SELECT cruise_key, COUNT(*) AS n FROM cruise
GROUP BY cruise_key HAVING COUNT(*) > 1"
)
if (nrow(cruise_dups) > 0) {
warning(glue("Duplicate cruise_key values: {nrow(cruise_dups)}"))
}
# cruise bridge coverage
bridge_coverage <- dbGetQuery(
con,
"SELECT
COUNT(*) AS total_casts,
SUM(CASE WHEN ship_key IS NOT NULL THEN 1 ELSE 0 END) AS with_ship_key,
SUM(CASE WHEN cruise_key IS NOT NULL THEN 1 ELSE 0 END) AS with_cruise_key
FROM casts"
)
bridge_coverage |> datatable(caption = "Cruise bridge coverage")
message("Cross-dataset validation complete")
```
### Re-export Casts Parquet
Update casts.parquet with the new ship_key and cruise_key columns.
```{r}
#| label: rewrite_casts_parquet
# re-export casts parquet (now includes ship_key, cruise_key)
casts_pqt <- file.path(dir_parquet, "casts.parquet")
cols <- DBI::dbGetQuery(
con,
glue(
"SELECT column_name FROM information_schema.columns
WHERE table_name = 'casts'"
)
)$column_name
prov_cols <- c("_source_file", "_source_row", "_source_uuid", "_ingested_at")
export_cols <- setdiff(cols, prov_cols)
DBI::dbExecute(
con,
glue(
"COPY (SELECT {paste(export_cols, collapse = ', ')} FROM casts)
TO '{casts_pqt}' (FORMAT PARQUET, COMPRESSION 'snappy')"
)
)
message("Re-exported casts.parquet with ship_key + cruise_key")
# export _new delta sidecars for modified dependency tables (from calcofi.modifies)
# these are NOT in the manifest — picked up by build_release_table_registry()
for (tbl in modifies_tables) {
pk_col <- modifies_pks[[tbl]]$pk_col
keys_old <- modifies_pks[[tbl]]$keys
keys_new <- dbGetQuery(con, glue("SELECT {pk_col} FROM {tbl}"))[[1]]
additions <- setdiff(keys_new, keys_old)
if (length(additions) > 0) {
new_tbl <- paste0(tbl, "_new")
pq_path <- file.path(dir_parquet, paste0(new_tbl, ".parquet"))
vals <- paste(shQuote(additions, "sh"), collapse = ", ")
export_parquet(
con,
glue("SELECT * FROM {tbl} WHERE {pk_col} IN ({vals})"),
pq_path
)
message(glue("{length(additions)} new {tbl} row(s) -> {new_tbl}.parquet"))
} else {
message(glue("No new {tbl} rows — {tbl}_new not exported"))
}
}
```
## Cleanup
```{r}
#| label: cleanup
# close local wrangling database connection
close_duckdb(con)
message("Local wrangling database connection closed")
# note: parquet outputs are in data/parquet/calcofi_bottle/
# these can be used to update the Working DuckLake in a separate workflow
message(glue("Parquet outputs written to: {dir_parquet}"))
message(glue("GCS outputs at: gs://calcofi-db/ingest/{dir_label}/"))
```
## TODO
- [x] Review column names and apply db naming conventions, per @docs/db.qmd and @workflows/README_PLAN.qmd: cast_cnt -> cast_id, btl_cnt -> bottle_id, sta_id -> site_key, etc.
- [x] Pivot longer casts/bottle tables for easier analysis. Created bottle_measurement (value/prec/qual) and cast_condition tables, plus measurement_type reference.
- [x] Consolidate casts temporal columns (date+time -> datetime_utc), drop 16 derivable columns, create casts_derived VIEW.
- [ ] Link possible fields to @workflows/ingest_swfsc_ichthyo.qmd. Flag mismatches.
- [x] Cruise bridge prep: renamed `Cruise_ID` → `cruise_key_0` (was `cruise_key`) to avoid collision with SWFSC `cruise_key` (YYMMKK). Merge workflow adds `cruise_key` as new column derived from `datetime_utc` + `ship_key`.
- [ ] Validate casts.ship_code -> ship.ship_nodc FK after integration with swfsc data.
- [ ] Review documentation for more comment descriptions at [Data \>
Data Formats \| CalCOFI.org](https://calcofi.com/index.php?option=com_content&view=category&id=73&Itemid=993)
::: {.callout-caution collapse="true"}
## Session Info
```{r session_info}
devtools::session_info()
```
:::