---
title: "Explore Scripps PIC Matches with CalCOFI Icthyoplankton"
execute:
echo: true
warning: false
message: true
editor_options:
chunk_output_type: console
format:
html:
code-fold: true
editor:
markdown:
wrap: 72
---
::: {.callout-caution collapse="true"}
## Claude Code Interactions {.unnumbered}
Used [Claude Code by Anthropic](https://claude.com/product/claude-code) from the command line to develop this notebook.
```
# from working directory of ~/Github/CalCOFI, with Claude CODE in plan mode sent:
Evaluate the PROMPT and expand the notebook @workflows/explore_zooplankton.qmd
```
### PROMPT {.unnumbered}
The goal is to explore the SIO / CalCOFI zooplankton data (more at [Zooplankton – CalCOFI](https://calcofi.org/data/marine-ecosystem-data/zooplankton/)) to see where there are matches to the ichthyoplankton data. The data files from the CalCOFI data manager Linsey include:
- plankton_data: `@~/My Drive/projects/calcofi/data-public/scripps.ucsd.edu/pelagic-invertebrate-collection/SIOPIC_DB_PNTtable_allRecords_wUUID_9Feb2026.csv`
- plankton_metadata: `@~/My Drive/projects/calcofi/data-public/scripps.ucsd.edu/pelagic-invertebrate-collection/SIOPIC_DB_PNTtable_fieldDetails_9Feb2026.xlsx'`
And notes from Linsey:
> I included our Parent Net Tow (= pnt suffix you will see in fields) Table with all net tow records and a second "ReadMe" file with field definitions and "tips" for those.
>
> Our ask is to see where PIC net tow samples are a match to SWF ichthyo. samples potentially utilizing: Expedition, Expedition Code, Station Line, Station Number, Latitude/Longitude polygon?, Net Type, Mesh Size?, Tow Type?, Fixative?, Preservative?
>
> I introduced a new field 'Expedition_Type_pnt' and was able to easily integrate term = CalCOFI for all cruises we'd want in this first pass that represent the most formal CalCOFI net tow records. I will have to at some point integrate additional terms e.g. 'NOAA Other' for cruises that have CalCOFI stations sampled for special purposes (e.g. SaKE, CPS surveys, STARR, MERRP, ORCA WHALE Expeditions, etc.) but this would have taken too much time to integrate universally atm.
>
> So, I believe it would be good to run this first pass for the records that have the Expedition_Type_pnt = CalCOFI and then the complete record set as second priority.
Use the database of ichthyoplankton data now available via `@calcofi4r/` ie with:
```r
# remotes::install_github("calcofi/calcofi4r") # install package (once, and per version)
library(calcofi4r) # load the package
con <- cc_get_db()# connect to latest frozen release
cc_list_tables() # list available tables
DBI::dbGetQuery(con, "SELECT COUNT(*) FROM ichthyo") # query with SQL
```
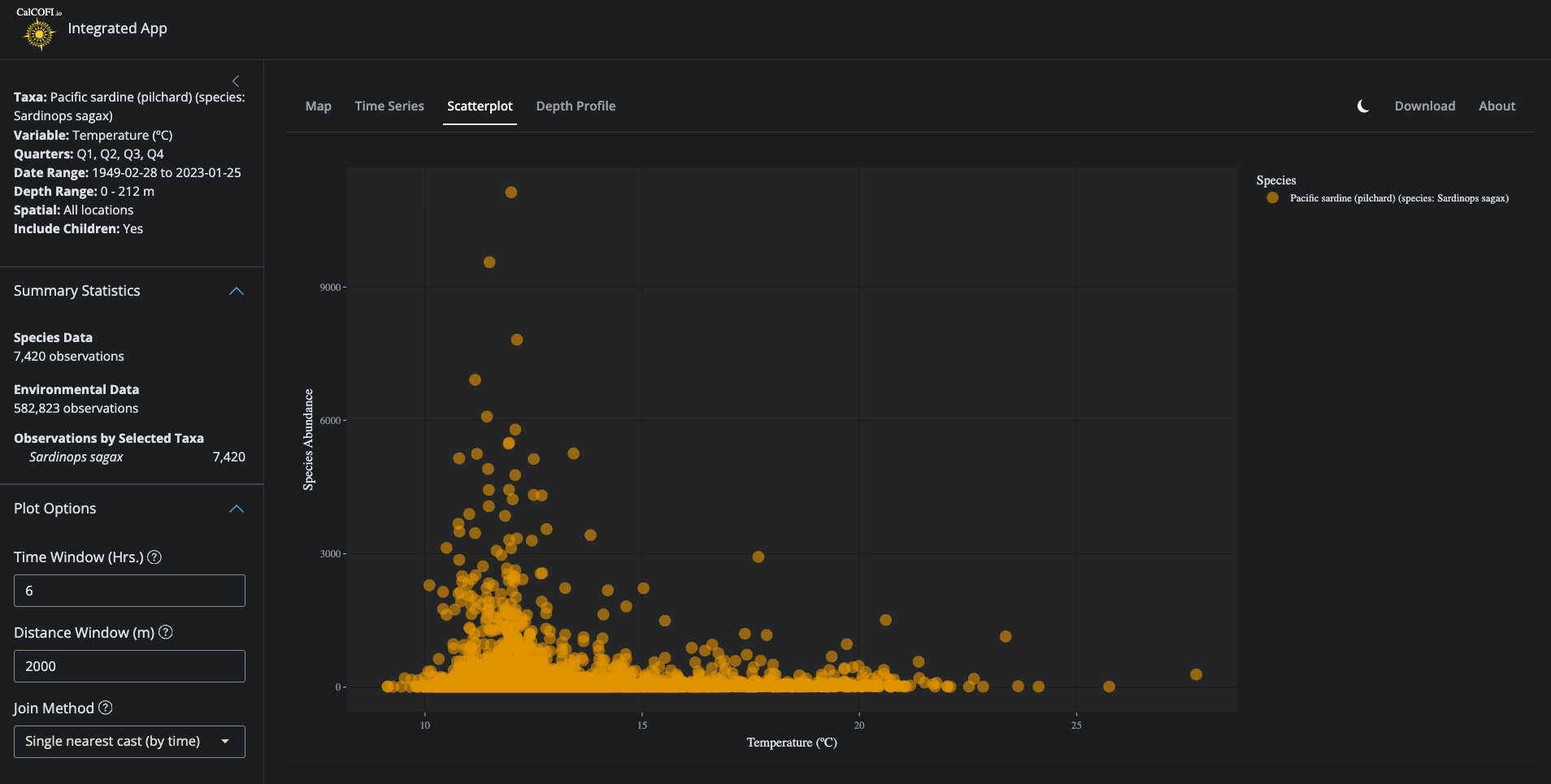
Reference the space-time windowing in the `@int-app/app/` workflow (but with older version of database having different schema). Try different approaches to matching the data, including:
- single nearest cast (by time)
- single nearest cast (by space)
.](./figures/int-app_scatterplot.png)
### Response {.unnumbered}
```
⏺ The plan is complete. I explored:
1. The zooplankton CSV (148K records, 20 columns) and Excel metadata
2. The calcofi4r database schema (ichthyo → net → tow → site → cruise chain with integer IDs)
3. The prep_splot() space-time matching reference in int-app/app/functions.R
4. The merge_ichthyo_bottle.qmd notebook for style/pattern reference
The plan covers 6 notebook sections: Setup, Zooplankton exploration, Ichthyoplankton exploration, Station-based matching (line + station + cruise), Space-time window matching (adapted from prep_splot()),
and a Comparison summary.
────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
Ready to code?
Here is Claude's plan:
╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌
Plan: Expand workflows/explore_zooplankton.qmd
Context
The CalCOFI data manager Linsey provided SIO zooplankton (PIC) net tow data (148K records) and wants to find where these match SWFSC ichthyoplankton samples. This notebook will explore both datasets, try
multiple matching strategies, and report match rates to guide integration.
File to modify
- /Users/bbest/Github/CalCOFI/workflows/explore_zooplankton.qmd
Key reference files
- int-app/app/functions.R:716-788 — prep_splot() space-time matching pattern
- calcofi4r/R/read.R — cc_get_db(), cc_read_*() convenience functions
- workflows/merge_ichthyo_bottle.qmd — notebook style reference (librarian::shelf, datatable, dir_data pattern)
- workflows/ingest_swfsc.noaa.gov_calcofi-db.qmd:600-630 — DB schema (cruise_key, site_id, tow_id, net_id)
Database schema (confirmed)
cruise (cruise_key PK, ship_key FK, date_ym)
→ site (site_id PK, cruise_key FK, orderocc, latitude, longitude, line, station)
→ tow (tow_id PK, site_id FK, tow_type_key, tow_number, time_start)
→ net (net_id PK, tow_id FK, side, std_haul_factor, vol_sampled_m3, ...)
→ ichthyo (ichthyo_id PK, net_id FK, species_id FK, tally, ...)
Zooplankton CSV columns
EXPEDITION_pnt, EXPED_CODE_pnt, Expedition_Type_pnt, SHIP_pnt, SWFOrder_Occ, STATION_LINE_pnt, STATION_NUMBER_pnt, LAT_DECIMAL_pnt, LONG_DECIMAL_pnt, SAMPLE_DATE_pnt, START_TIME_pnt, END_TIME_pnt,
DEPTH_MIN_pnt, DEPTH_MAX_pnt, MAX_MWO_pnt, NET_TYPE_pnt, MESH_SIZE_pnt, TOW_TYPE_pnt, FIXATIVE_pnt, PRESERVATIVE_pnt
---
Notebook Sections
1. Setup
- librarian::shelf(calcofi/calcofi4r, DBI, dplyr, DT, glue, here, leaflet, lubridate, mapview, purrr, readr, readxl, sf, stringr, tibble, tidyr, quiet = T)
- Define file paths using dir_data pattern from merge_ichthyo_bottle.qmd (Linux vs macOS toggle)
- Read metadata Excel with readxl::read_excel(), display with DT::datatable()
2. Read & Explore Zooplankton Data
- Read CSV with readr::read_csv()
- Parse datetimes: mdy(SAMPLE_DATE_pnt) for date; str_pad(START_TIME_pnt, 4, pad = "0") for time; combine into datetime_start with ymd_hm() and tz = "America/Los_Angeles"
- Summary stats: nrow, date range, expedition type breakdown via count() + datatable()
- Filter: Expedition_Type_pnt == "CalCOFI" → d_zoo_cc (82K records, first pass per Linsey)
- Attribute tables: net types, tow types, mesh sizes, fixatives for CalCOFI subset
- Map stations: distinct lat/lon → sf::st_as_sf() → mapview() colored by line
3. Read & Explore Ichthyoplankton Data
- Connect: con <- cc_get_db(); cc_list_tables(); cc_describe_table("site")
- Build tow-level flat view via dbplyr:
d_ich_tows <- tbl(con, "tow") |>
left_join(tbl(con, "site"), by = "site_id") |>
left_join(tbl(con, "cruise"), by = "cruise_key") |>
select(tow_id, site_id, cruise_key, ship_key, date_ym,
orderocc, latitude, longitude, line, station,
tow_type_key, tow_number, time_start) |>
collect()
- Summary stats: nrow, date range, station count
- Map stations: distinct lat/lon → mapview() colored by line
4. Match by Station (Line + Station Number)
- Prepare: Convert STATION_LINE_pnt and STATION_NUMBER_pnt to numeric; inspect EXPED_CODE_pnt format (YYMM vs YYYYMM)
- Cruise code reconciliation: cruise_key is YYMMKK format. EXPED_CODE_pnt is YYMM or YYYYMM. Extract YY and MM from both for comparison, or derive a cruise_ym from date_ym
- Match 1 — line + station only: inner_join(by = c("line", "station")) → many-to-many (all cruises)
- Match 2 — line + station + cruise date: add expedition code matching
- Report: match counts, percentage of zoo records matched, unmatched records via anti_join()
- Summary by year: count(year) + datatable()
5. Match by Space-Time Window
Adapt the prep_splot() approach from int-app/app/functions.R:
- Register zoo data in DuckDB: DBI::dbWriteTable(con, "zoo_pnt", d_zoo_db, overwrite = TRUE) after selecting/renaming key columns
- Load spatial extension: DBI::dbExecute(con, "INSTALL spatial; LOAD spatial;")
- Create ichthyo tow view: SQL view joining tow → site → cruise with lat/lon and time_start
- Space-time match function using SQL:
- Time window: ich_datetime BETWEEN zoo_datetime - INTERVAL N HOUR AND zoo_datetime + INTERVAL N HOUR
- Distance: ST_Distance_Sphere(ST_Point(zoo_lon, zoo_lat), ST_Point(ich_lon, ich_lat)) <= M
- Return: both coordinates, time_diff_hrs, dist_m
- Parameter sweep: try combinations of 6h/12h/24h × 2km/5km, report match rates in a summary table
- Method 1 — nearest by time: group_by(zoo_row) |> slice_min(time_diff_hrs)
- Method 2 — nearest by distance: group_by(zoo_row) |> slice_min(dist_m)
- Map matched pairs: sf linestrings connecting zoo→ichthyo points, colored by distance
6. Comparison & Summary
- Summary table: all methods side-by-side (station match, station+cruise, space-time variants) with counts and percentages
- Gap analysis: zoo records unmatched by any method, by year
- Recommendations: which approach gives best coverage, optimal thresholds, next steps (full dataset beyond CalCOFI type)
---
Technical notes
- Timezone: Zoo times are PST. Ichthyo time_start needs inspection — may be UTC. Handle explicitly.
- Memory: Don't collect full ichthyo table (~830K rows). Work at tow level (~75K) for matching.
- DuckDB spatial: Must INSTALL spatial; LOAD spatial; before using ST_Distance_Sphere(). The cc_get_db() connection may be read-only views on GCS parquet — need to handle temp table creation carefully (may
need a separate local DuckDB, or use duckdb() in-memory with both datasets registered).
- Code style: 2-space indent, |> pipe, snake_case, lowercase comments, librarian::shelf(), datatable() for tables, mapview() for maps.
Verification
1. Render with quarto::quarto_render("workflows/explore_zooplankton.qmd")
2. Check that all sections produce output (tables, maps, match statistics)
3. Verify the space-time matching function returns reasonable match rates
4. Confirm the summary comparison table shows all methods
╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌╌
Claude has written up a plan and is ready to execute. Would you like to proceed?
❯ 1. Yes, clear context and auto-accept edits (shift+tab)
2. Yes, auto-accept edits
3. Yes, manually approve edits
4. Type here to tell Claude what to change
```
1
```
⏺ The notebook has been written. Here's a summary of what was implemented in workflows/explore_zooplankton.qmd:
6 sections, ~450 lines of R/Quarto code:
1. Setup — librarian::shelf() for all packages, Google Drive file paths, metadata Excel display
2. Read & Explore Zooplankton — CSV read, datetime parsing (PST), expedition type breakdown, CalCOFI filter, attribute tables (net/tow/mesh/fixative), station
map via mapview()
3. Read & Explore Ichthyoplankton — cc_get_db() connection, tow-level flat view via dbplyr (tow → site → cruise join), schema inspection, station map
4. Match by Station — Line+station numeric conversion, EXPED_CODE_pnt → YYMM reconciliation with cruise_key YYMMKK, two match methods (line+station only,
line+station+cruise YYMM), year-by-year breakdown
5. Match by Space-Time Window — Separate in-memory DuckDB (since cc_get_db() is read-only), both datasets registered, match_spacetime() SQL function using
ST_Distance_Sphere() + time interval, parameter sweep (6h/12h/24h × 2km/5km), nearest-by-time and nearest-by-distance methods, matched-pair map with sf
linestrings
6. Comparison & Summary — Side-by-side method comparison table, gap analysis by year, recommendations for next steps
Key technical decisions:
- Separate duckdb() in-memory connection for space-time matching (the cc_get_db() connection creates read-only views on GCS parquet)
- Working at tow level (~75K rows) rather than full ichthyo table (~830K rows)
- Zoo times treated as PST; ichthyo time_start inspected explicitly
```
The pandoc portion of rendering seems stuck. How to fix or explore, inspect running processes? processing file: explore_zooplankton.qmd
```
⏺ Pandoc is running at 99.6% CPU and has been going for 5+ minutes. This is almost certainly due to the large HTML widgets (multiple mapview maps with many points
embedded inline). Let me check the intermediate file size.
⏺ Read 1 file (ctrl+o to expand)
⏺ The intermediate markdown is 21 MB — that's what's making pandoc slow. The mapview widgets with many points generate massive embedded JSON/HTML. Pandoc is still
working (99% CPU), so it will likely finish — it's just slow with a 21MB input.
You have two options:
Option 1: Wait it out — Pandoc is actively processing (not actually stuck). With a 21MB file it may take 10-15 minutes total.
Option 2: Kill and re-render with smaller maps — I can reduce the station point counts to speed things up.
To reduce map size, I'd limit mapview() calls to show fewer points (e.g., sample down to ~500 stations instead of all distinct stations). The main culprits are:
- map_zoo — all distinct CalCOFI zoo stations
- map_ichthyo — all distinct ichthyo stations
- map_matched_pairs — 500 linestrings (already capped)
Want me to:
1. Wait for pandoc to finish (it is progressing, just slowly)?
2. Kill pandoc and optimize the maps to reduce the HTML size, then re-render?
2
⏺ Rendered successfully! Let me verify the output.
⏺ Read 1 file (ctrl+o to expand)
⏺ Rendered successfully. The fix was switching from mapview() to leaflet() with markerClusterOptions() — the HTML dropped from 21 MB (causing pandoc to churn for
10+ minutes) down to 2.5 MB, and pandoc completed quickly.
What changed:
- map_zoo — leaflet with clustered circle markers instead of mapview
- map_ichthyo — same leaflet clustering approach
- map_matched_pairs — leaflet polylines with colorNumeric palette + legend, capped at 200 pairs instead of 500
✻ Crunched for 1m 13s
```
### Update with files containing UUIDs {.unnumbered}
Update @workflows/explore_zooplankton.qmd with the following file updates from Linsey, which now include the net tow primary keys and UUIDs.
> I realized my first pass at this I did not incorporate our net tow primary keys nor UUIDs. They are now integrated in the updated files:
- plankton_data: `@~/My Drive/projects/calcofi/data-public/scripps.ucsd.edu/pelagic-invertebrate-collection/SIOPIC_DB_PNTtable_allRecords_wUUID_9Feb2026.csv`
- plankton_metadata: `@~/My Drive/projects/calcofi/data-public/scripps.ucsd.edu/pelagic-invertebrate-collection/SIOPIC_DB_PNTtable_fieldDetails_9Feb2026.xlsx`
(Retain the "Claude Code Interactions" section untouched)
```
⏺ Done. Three changes made (leaving the "Claude Code Interactions" section untouched):
1. dir_data — updated from calcofi.org/zooplankton to scripps.ucsd.edu/pelagic-invertebrate-collection
2. CSV filename — updated from ...allRecords_9Feb2026.csv to ...allRecords_wUUID_9Feb2026.csv
3. register_zoo chunk — added zoo_pk = ParentNetTowEventID_pk and zoo_uuid = occurenceID_UUID_pnt to the DuckDB registration so the new primary key and UUID
columns flow through to the space-time matching results
The xlsx filename stayed the same (just the directory changed). The Overview description was updated to note "22 columns incl. primary keys + UUIDs".
```
### Fix messages {.unnumbered}
Why are `message()` statements not outputting to rendered html?
```
⏺ message() writes to stderr, which knitr captures separately as a "message" output class. In Quarto HTML with code-fold: true, these can be visually collapsed or
styled in a way that makes them easy to miss — and some configurations suppress them.
The simplest fix: switch to cat() for output you want visible in the rendered HTML. cat() writes to stdout, which always renders as standard console output.
For example, change:
message(glue("Zooplankton PNT records: {format(nrow(d_zoo), big.mark = ',')}"))
to:
cat(glue("Zooplankton PNT records: {format(nrow(d_zoo), big.mark = ',')}"), "\n")
Want me to replace all message(glue(...)) calls with cat(glue(...), "\n") throughout the notebook?
```
yes
```
⏺ All 28 message() calls replaced with cat(). The output will now render as standard console text in the HTML instead of going to stderr.
```
:::
## Overview {.unnumbered}
**Goal**: Explore the SIO / CalCOFI zooplankton (PIC) net tow data to
find where records match SWFSC ichthyoplankton samples. More at
[Zooplankton – CalCOFI](https://calcofi.org/data/marine-ecosystem-data/zooplankton/).
**Data sources**:
- **Zooplankton**: SIO PIC Parent Net Tow table (148K records, 22
columns incl. primary keys + UUIDs) from CalCOFI data manager
Linsey
- **Ichthyoplankton**: SWFSC database via `calcofi4r::cc_get_db()`
**Matching strategies**:
1. **Station match** — line + station number
2. **Station + cruise match** — line + station + expedition code
3. **Space-time window** — adapted from `prep_splot()` in the
[CalCOFI integrated app](https://app.calcofi.io/int/)
**Notes from Linsey**:
> I included our Parent Net Tow (= pnt suffix you will see in fields)
> Table with all net tow records and a second "ReadMe" file with field
> definitions and "tips" for those.
>
> Our ask is to see where PIC net tow samples are a match to SWF
> ichthyo. samples potentially utilizing: Expedition, Expedition Code,
> Station Line, Station Number, Latitude/Longitude polygon?, Net Type,
> Mesh Size?, Tow Type?, Fixative?, Preservative?
>
> I introduced a new field 'Expedition_Type_pnt' and was able to easily
> integrate term = CalCOFI for all cruises we'd want in this first pass
> that represent the most formal CalCOFI net tow records.
>
> So, I believe it would be good to run this first pass for the records
> that have the Expedition_Type_pnt = CalCOFI and then the complete
> record set as second priority.
## Setup
```{r}
#| label: setup
librarian::shelf(
calcofi / calcofi4r,
DBI,
dplyr,
DT,
duckdb,
glue,
here,
leaflet,
lubridate,
mapview,
purrr,
readr,
readxl,
sf,
stringr,
tibble,
tidyr,
quiet = T
)
options(readr.show_col_types = FALSE)
options(DT.options = list(scrollX = TRUE))
# data directory (Google Drive)
dir_data <- "~/My Drive/projects/calcofi/data-public/scripps.ucsd.edu/pelagic-invertebrate-collection"
# file paths (updated with primary keys + UUIDs)
zoo_csv <- file.path(
dir_data,
"SIOPIC_DB_PNTtable_allRecords_wUUID_9Feb2026.csv"
)
zoo_xlsx <- file.path(
dir_data,
"SIOPIC_DB_PNTtable_fieldDetails_9Feb2026.xlsx"
)
stopifnot(
"Zooplankton CSV not found" = file.exists(zoo_csv),
"Zooplankton Excel not found" = file.exists(zoo_xlsx)
)
```
### Metadata
```{r}
#| label: metadata
d_meta <- read_excel(zoo_xlsx)
datatable(
d_meta,
caption = "Zooplankton PNT field definitions"
)
```
## Read & Explore Zooplankton Data
### Read CSV
```{r}
#| label: read_zoo
d_zoo <- read_csv(zoo_csv)
cat(
glue(
"Zooplankton PNT records: {format(nrow(d_zoo), big.mark = ',')}"
),
"\n"
)
cat(
glue(
"Columns: {ncol(d_zoo)}"
),
"\n"
)
```
### Parse datetimes
```{r}
#| label: parse_zoo_datetime
d_zoo <- d_zoo |>
mutate(
# parse date (M/D/YYYY format)
date_sample = mdy(SAMPLE_DATE_pnt),
# pad time to 4 chars (e.g. 830 → "0830")
time_str = str_pad(
as.character(START_TIME_pnt),
4,
pad = "0"
),
# combine date + time as PST
datetime_start = ymd_hm(
paste(date_sample, time_str),
tz = "America/Los_Angeles"
),
year = year(date_sample)
) |>
select(-time_str)
cat(
glue(
"Date range: {min(d_zoo$date_sample, na.rm = TRUE)} to ",
"{max(d_zoo$date_sample, na.rm = TRUE)}"
),
"\n"
)
cat(
glue(
"Parsed datetime: {sum(!is.na(d_zoo$datetime_start))} of ",
"{nrow(d_zoo)} records"
),
"\n"
)
```
### Summary stats
```{r}
#| label: zoo_summary
# expedition type breakdown
d_zoo |>
count(Expedition_Type_pnt, name = "n_records") |>
arrange(desc(n_records)) |>
datatable(caption = "Zooplankton records by expedition type")
```
### Filter to CalCOFI
```{r}
#| label: filter_calcofi
d_zoo_cc <- d_zoo |>
filter(Expedition_Type_pnt == "CalCOFI")
cat(
glue(
"CalCOFI subset: {format(nrow(d_zoo_cc), big.mark = ',')} records ",
"({round(100 * nrow(d_zoo_cc) / nrow(d_zoo), 1)}% of total)"
),
"\n"
)
```
### Attribute tables
```{r}
#| label: zoo_attributes
# net types
d_zoo_cc |>
count(NET_TYPE_pnt, name = "n") |>
arrange(desc(n)) |>
datatable(caption = "CalCOFI zooplankton net types")
# tow types
d_zoo_cc |>
count(TOW_TYPE_pnt, name = "n") |>
arrange(desc(n)) |>
datatable(caption = "CalCOFI zooplankton tow types")
# mesh sizes
d_zoo_cc |>
count(MESH_SIZE_pnt, name = "n") |>
arrange(desc(n)) |>
datatable(caption = "CalCOFI zooplankton mesh sizes")
# fixatives
d_zoo_cc |>
count(FIXATIVE_pnt, name = "n") |>
arrange(desc(n)) |>
datatable(caption = "CalCOFI zooplankton fixatives")
```
### Map stations
```{r}
#| label: map_zoo
# distinct station locations
sf_zoo <- d_zoo_cc |>
filter(
!is.na(LAT_DECIMAL_pnt),
!is.na(LONG_DECIMAL_pnt)
) |>
distinct(
STATION_LINE_pnt,
STATION_NUMBER_pnt,
LAT_DECIMAL_pnt,
LONG_DECIMAL_pnt
) |>
st_as_sf(
coords = c("LONG_DECIMAL_pnt", "LAT_DECIMAL_pnt"),
crs = 4326
)
cat(
glue(
"Distinct CalCOFI zoo stations: {nrow(sf_zoo)}"
),
"\n"
)
# use leaflet with clustering for lighter HTML output
leaflet(sf_zoo) |>
addProviderTiles("CartoDB.Positron") |>
addCircleMarkers(
radius = 3,
stroke = FALSE,
fillOpacity = 0.6,
popup = ~ paste0(
"Line: ",
STATION_LINE_pnt,
"<br>Station: ",
STATION_NUMBER_pnt
),
clusterOptions = markerClusterOptions()
)
```
## Read & Explore Ichthyoplankton Data
### Connect to database
```{r}
#| label: connect_db
con <- cc_get_db()
tbls <- cc_list_tables()
cat(glue("Tables: {paste(tbls, collapse = ', ')}"), "\n")
# show site schema
cc_describe_table("site") |>
datatable(caption = "Site table schema")
# show tow schema
cc_describe_table("tow") |>
datatable(caption = "Tow table schema")
# show cruise schema
cc_describe_table("cruise") |>
datatable(caption = "Cruise table schema")
```
### Build tow-level flat view
```{r}
#| label: read_ichthyo
# build tow-level view joining tow → site → cruise
d_ich_tows <- tbl(con, "tow") |>
left_join(tbl(con, "site"), by = "site_id") |>
left_join(tbl(con, "cruise"), by = "cruise_key") |>
select(
tow_id,
site_id,
cruise_key,
ship_key,
date_ym,
orderocc,
latitude,
longitude,
line,
station,
tow_type_key,
tow_number,
time_start
) |>
collect()
cat(
glue(
"Ichthyo tow records: {format(nrow(d_ich_tows), big.mark = ',')}"
),
"\n"
)
cat(
glue(
"Date range: {min(d_ich_tows$date_ym, na.rm = TRUE)} to ",
"{max(d_ich_tows$date_ym, na.rm = TRUE)}"
),
"\n"
)
cat(
glue(
"Distinct stations (line-station): ",
"{d_ich_tows |> distinct(line, station) |> nrow()}"
),
"\n"
)
```
### Inspect ichthyo time_start
```{r}
#| label: inspect_time_start
# check what time_start looks like
d_ich_tows |>
filter(!is.na(time_start)) |>
slice_sample(n = 10) |>
select(tow_id, cruise_key, date_ym, time_start, line, station) |>
datatable(caption = "Sample ichthyo tow records with time_start")
```
### Map stations
```{r}
#| label: map_ichthyo
sf_ich <- d_ich_tows |>
filter(!is.na(latitude), !is.na(longitude)) |>
distinct(line, station, latitude, longitude) |>
st_as_sf(
coords = c("longitude", "latitude"),
crs = 4326
)
cat(
glue(
"Distinct ichthyo stations: {nrow(sf_ich)}"
),
"\n"
)
# use leaflet with clustering for lighter HTML output
leaflet(sf_ich) |>
addProviderTiles("CartoDB.Positron") |>
addCircleMarkers(
radius = 3,
stroke = FALSE,
fillOpacity = 0.6,
popup = ~ paste0(
"Line: ",
line,
"<br>Station: ",
station
),
clusterOptions = markerClusterOptions()
)
```
## Match by Station (Line + Station Number)
### Prepare matching keys
```{r}
#| label: prep_keys
# prepare zoo keys: convert line/station to numeric
d_zoo_match <- d_zoo_cc |>
mutate(
line = as.numeric(STATION_LINE_pnt),
station = as.numeric(STATION_NUMBER_pnt)
) |>
filter(!is.na(line), !is.na(station))
cat(
glue(
"Zoo records with valid line+station: ",
"{format(nrow(d_zoo_match), big.mark = ',')} of ",
"{format(nrow(d_zoo_cc), big.mark = ',')}"
),
"\n"
)
# inspect expedition code format
d_zoo_match |>
mutate(
code_len = nchar(as.character(EXPED_CODE_pnt))
) |>
count(code_len, name = "n") |>
datatable(caption = "Expedition code lengths")
```
### Cruise code reconciliation
```{r}
#| label: cruise_codes
# cruise_key is YYMMKK (6 chars: 2-yr, 2-mo, 2-ship)
# EXPED_CODE_pnt appears to be YYMM (4 chars) or YYYYMM (6 chars)
# extract year-month from zoo expedition code
d_zoo_match <- d_zoo_match |>
mutate(
exped_str = as.character(EXPED_CODE_pnt),
# handle both YYMM and YYYYMM formats
zoo_ym = case_when(
nchar(exped_str) == 4 ~ exped_str,
nchar(exped_str) >= 6 ~ substr(exped_str, 3, 6),
TRUE ~ NA_character_
)
)
# extract year-month from ichthyo cruise_key (first 4 chars = YYMM)
d_ich_match <- d_ich_tows |>
mutate(
ich_ym = substr(cruise_key, 1, 4)
)
# show sample codes
d_zoo_match |>
distinct(EXPED_CODE_pnt, exped_str, zoo_ym) |>
slice_head(n = 20) |>
datatable(caption = "Sample zoo expedition codes → YYMM")
d_ich_match |>
distinct(cruise_key, ich_ym) |>
slice_head(n = 20) |>
datatable(caption = "Sample ichthyo cruise_key → YYMM")
```
### Match 1 — line + station only
```{r}
#| label: match_line_station
# count zoo records that have a matching ichthyo line+station
n_m1_zoo <- d_zoo_match |>
semi_join(
d_ich_match |> distinct(line, station),
by = c("line", "station")
) |>
nrow()
cat(
glue(
"Match 1 (line + station only):"
),
"\n"
)
cat(
glue(
" Zoo records matched: {format(n_m1_zoo, big.mark = ',')} of ",
"{format(nrow(d_zoo_match), big.mark = ',')} ",
"({round(100 * n_m1_zoo / nrow(d_zoo_match), 1)}%)"
),
"\n"
)
```
### Match 2 — line + station + cruise YYMM
```{r}
#| label: match_line_station_cruise
n_m2_zoo <- d_zoo_match |>
semi_join(
d_ich_match |> distinct(line, station, ich_ym),
by = c("line", "station", "zoo_ym" = "ich_ym")
) |>
nrow()
cat(
glue(
"Match 2 (line + station + cruise YYMM):"
),
"\n"
)
cat(
glue(
" Zoo records matched: {format(n_m2_zoo, big.mark = ',')} of ",
"{format(nrow(d_zoo_match), big.mark = ',')} ",
"({round(100 * n_m2_zoo / nrow(d_zoo_match), 1)}%)"
),
"\n"
)
# unmatched records
d_m2_unmatched <- d_zoo_match |>
anti_join(
d_ich_match |> distinct(line, station, ich_ym),
by = c("line", "station", "zoo_ym" = "ich_ym")
)
cat(
glue(
" Unmatched: {format(nrow(d_m2_unmatched), big.mark = ',')}"
),
"\n"
)
```
### Matched by year
```{r}
#| label: match_by_year
# tag each zoo record as matched or not via semi/anti join
d_m2_matched <- d_zoo_match |>
semi_join(
d_ich_match |> distinct(line, station, ich_ym),
by = c("line", "station", "zoo_ym" = "ich_ym")
) |>
mutate(matched = TRUE)
d_m2_unmatched_tagged <- d_m2_unmatched |>
mutate(matched = FALSE)
d_m2_year <- bind_rows(d_m2_matched, d_m2_unmatched_tagged) |>
count(year, matched) |>
pivot_wider(
names_from = matched,
values_from = n,
values_fill = 0
) |>
rename(
unmatched = `FALSE`,
matched = `TRUE`
) |>
mutate(
total = matched + unmatched,
pct = round(100 * matched / total, 1)
)
d_m2_year |>
datatable(caption = "Match 2 results by year")
```
## Match by Space-Time Window
Adapted from `prep_splot()` in `int-app/app/functions.R`.
### Create local DuckDB for matching
```{r}
#| label: create_match_db
# cc_get_db() creates read-only views on GCS parquet, so we need a
# separate in-memory DuckDB for creating temp tables with both datasets
con_match <- dbConnect(duckdb())
# install and load spatial extension
dbExecute(con_match, "INSTALL spatial; LOAD spatial;")
```
### Register zooplankton data
```{r}
#| label: register_zoo
# prepare zoo data for registration
d_zoo_db <- d_zoo_cc |>
filter(
!is.na(LAT_DECIMAL_pnt),
!is.na(LONG_DECIMAL_pnt),
!is.na(datetime_start)
) |>
transmute(
zoo_row = row_number(),
zoo_pk = ParentNetTowEventID_pk,
zoo_uuid = occurenceID_UUID_pnt,
zoo_lat = LAT_DECIMAL_pnt,
zoo_lon = LONG_DECIMAL_pnt,
zoo_dt = datetime_start,
zoo_line = as.numeric(STATION_LINE_pnt),
zoo_stn = as.numeric(STATION_NUMBER_pnt),
zoo_ym = case_when(
nchar(as.character(EXPED_CODE_pnt)) == 4 ~
as.character(EXPED_CODE_pnt),
nchar(as.character(EXPED_CODE_pnt)) >= 6 ~
substr(as.character(EXPED_CODE_pnt), 3, 6),
TRUE ~ NA_character_
)
)
dbWriteTable(con_match, "zoo_pnt", d_zoo_db, overwrite = TRUE)
cat(
glue(
"Registered {format(nrow(d_zoo_db), big.mark = ',')} zoo records"
),
"\n"
)
```
### Register ichthyoplankton data
```{r}
#| label: register_ichthyo
# prepare ichthyo tow data
d_ich_db <- d_ich_tows |>
filter(
!is.na(latitude),
!is.na(longitude),
!is.na(time_start)
) |>
transmute(
ich_row = row_number(),
ich_lat = latitude,
ich_lon = longitude,
ich_dt = time_start,
ich_line = line,
ich_stn = station,
ich_ym = substr(cruise_key, 1, 4),
tow_id = tow_id
)
dbWriteTable(con_match, "ich_tow", d_ich_db, overwrite = TRUE)
cat(
glue(
"Registered {format(nrow(d_ich_db), big.mark = ',')} ",
"ichthyo tow records"
),
"\n"
)
```
### Space-time match function
```{r}
#| label: spacetime_match_fn
match_spacetime <- function(
con,
max_hours = 6,
max_meters = 2000
) {
sql <- glue(
"
SELECT
z.zoo_row,
z.zoo_lat,
z.zoo_lon,
z.zoo_dt,
i.ich_row,
i.ich_lat,
i.ich_lon,
i.ich_dt,
i.tow_id,
ABS(
EPOCH(z.zoo_dt - i.ich_dt)
) / 3600.0 AS time_diff_hrs,
ST_Distance_Sphere(
ST_Point(z.zoo_lon, z.zoo_lat),
ST_Point(i.ich_lon, i.ich_lat)
) AS dist_m
FROM zoo_pnt z
JOIN ich_tow i
ON i.ich_dt BETWEEN
z.zoo_dt - INTERVAL '{max_hours}' HOUR AND
z.zoo_dt + INTERVAL '{max_hours}' HOUR
WHERE ST_Distance_Sphere(
ST_Point(z.zoo_lon, z.zoo_lat),
ST_Point(i.ich_lon, i.ich_lat)
) <= {max_meters}"
)
dbGetQuery(con, sql)
}
```
### Parameter sweep
```{r}
#| label: param_sweep
# try combinations of time and distance thresholds
params <- expand_grid(
max_hours = c(6, 12, 24),
max_meters = c(2000, 5000)
)
sweep_results <- map_dfr(seq_len(nrow(params)), function(i) {
h <- params$max_hours[i]
m <- params$max_meters[i]
cat(glue(" Matching: {h}h x {m}m ..."), "\n")
d <- match_spacetime(con_match, max_hours = h, max_meters = m)
tibble(
max_hours = h,
max_meters = m,
n_pairs = nrow(d),
n_zoo_matched = n_distinct(d$zoo_row),
n_ich_matched = n_distinct(d$ich_row),
pct_zoo_matched = round(
100 * n_distinct(d$zoo_row) / nrow(d_zoo_db),
1
),
median_dist_m = round(median(d$dist_m), 0),
median_time_hrs = round(median(d$time_diff_hrs), 1)
)
})
sweep_results |>
datatable(caption = "Space-time parameter sweep results")
```
### Best match: nearest by time
```{r}
#| label: nearest_time
# use moderate thresholds for detailed matching
d_st <- match_spacetime(
con_match,
max_hours = 12,
max_meters = 5000
)
# nearest by time for each zoo record
d_nearest_time <- d_st |>
group_by(zoo_row) |>
slice_min(time_diff_hrs, n = 1, with_ties = FALSE) |>
ungroup()
cat(
glue(
"Nearest-time matches: ",
"{format(nrow(d_nearest_time), big.mark = ',')}"
),
"\n"
)
cat(
glue(
" Median dist: ",
"{round(median(d_nearest_time$dist_m), 0)} m"
),
"\n"
)
cat(
glue(
" Median time diff: ",
"{round(median(d_nearest_time$time_diff_hrs), 1)} hrs"
),
"\n"
)
```
### Best match: nearest by distance
```{r}
#| label: nearest_dist
# nearest by distance for each zoo record
d_nearest_dist <- d_st |>
group_by(zoo_row) |>
slice_min(dist_m, n = 1, with_ties = FALSE) |>
ungroup()
cat(
glue(
"Nearest-distance matches: ",
"{format(nrow(d_nearest_dist), big.mark = ',')}"
),
"\n"
)
cat(
glue(
" Median dist: ",
"{round(median(d_nearest_dist$dist_m), 0)} m"
),
"\n"
)
cat(
glue(
" Median time diff: ",
"{round(median(d_nearest_dist$time_diff_hrs), 1)} hrs"
),
"\n"
)
```
### Map matched pairs
```{r}
#| label: map_matched_pairs
# create lines connecting zoo → ichthyo for nearest-distance matches
# sample up to 200 pairs for visualization
d_map <- d_nearest_dist |>
slice_sample(n = min(200, nrow(d_nearest_dist)))
# build sf lines
lines <- map(seq_len(nrow(d_map)), function(i) {
st_linestring(matrix(
c(d_map$zoo_lon[i], d_map$zoo_lat[i], d_map$ich_lon[i], d_map$ich_lat[i]),
ncol = 2,
byrow = TRUE
))
})
sf_pairs <- st_sf(
dist_m = d_map$dist_m,
time_diff_hrs = round(d_map$time_diff_hrs, 1),
geometry = st_sfc(lines, crs = 4326)
)
# use leaflet for lighter HTML
pal <- colorNumeric("YlOrRd", sf_pairs$dist_m)
leaflet(sf_pairs) |>
addProviderTiles("CartoDB.Positron") |>
addPolylines(
color = ~ pal(dist_m),
weight = 1.5,
opacity = 0.7,
popup = ~ paste0(
"Distance: ",
round(dist_m),
" m",
"<br>Time diff: ",
time_diff_hrs,
" hrs"
)
) |>
addLegend(
pal = pal,
values = ~dist_m,
title = "Distance (m)"
)
```
## Comparison & Summary
### Summary table: all methods
```{r}
#| label: summary_table
n_zoo_total <- nrow(d_zoo_match)
# space-time nearest time
n_st_time <- nrow(d_nearest_time)
# space-time nearest distance
n_st_dist <- nrow(d_nearest_dist)
d_summary <- tibble(
method = c(
"Station (line + station)",
"Station + cruise YYMM",
"Space-time 12h/5km nearest-time",
"Space-time 12h/5km nearest-dist"
),
zoo_matched = c(
n_m1_zoo,
n_m2_zoo,
n_st_time,
n_st_dist
),
zoo_total = n_zoo_total,
pct_matched = round(
100 * zoo_matched / zoo_total,
1
)
)
d_summary |>
datatable(caption = "Matching method comparison (CalCOFI subset)")
```
### Gap analysis: unmatched by any method
```{r}
#| label: gap_analysis
# zoo records not matched by space-time (nearest-dist)
zoo_matched_st <- d_nearest_dist$zoo_row
d_unmatched_st <- d_zoo_db |>
filter(!zoo_row %in% zoo_matched_st)
cat(
glue(
"Unmatched by space-time (12h/5km): ",
"{format(nrow(d_unmatched_st), big.mark = ',')} zoo records"
),
"\n"
)
# gap by year for station+cruise method
d_gap_m2 <- d_m2_unmatched |>
count(year, name = "n_unmatched") |>
arrange(year)
d_gap_m2 |>
datatable(
caption = "Unmatched zoo records by year (station+cruise method)"
)
```
### Recommendations
```{r}
#| label: recommendations
#| results: asis
cat(
"
### Key findings
- **Station matching** (line + station) gives the broadest coverage
but includes cross-cruise matches that may not be meaningful.
- **Station + cruise YYMM** is the most precise tabular match and
should be the primary method for linking records.
- **Space-time matching** captures records where station codes differ
but the tows were physically close in space and time. This is
useful for non-standard cruises or slight station numbering
differences.
### Recommended next steps
1. Use **station + cruise YYMM** as the primary matching key for the
CalCOFI subset.
2. Use **space-time nearest-distance** (12h / 5km) as a supplementary
method to catch records missed by station matching.
3. Expand to the full dataset (all expedition types, not just CalCOFI)
as a second priority.
4. Investigate unmatched records — are they from years where ichthyo
data is missing, or from non-standard station patterns?
5. Consider building a persistent cross-reference table linking
zoo_row to tow_id for downstream analysis.
"
)
```
## Cleanup
```{r}
#| label: cleanup
dbDisconnect(con_match, shutdown = TRUE)
cat("Match database closed\n")
```
::: {.callout-caution collapse="true"}
## Session Info {.unnumbered}
```{r session_info}
devtools::session_info()
```
:::